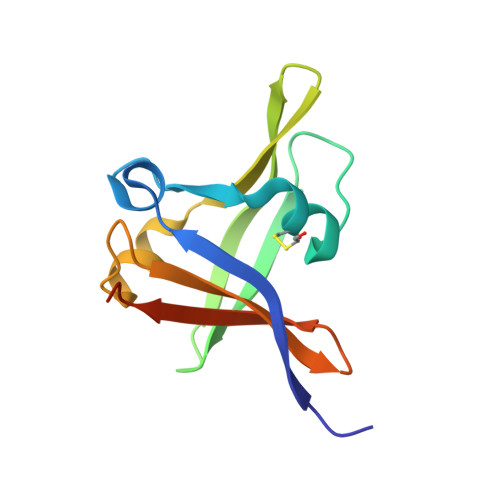
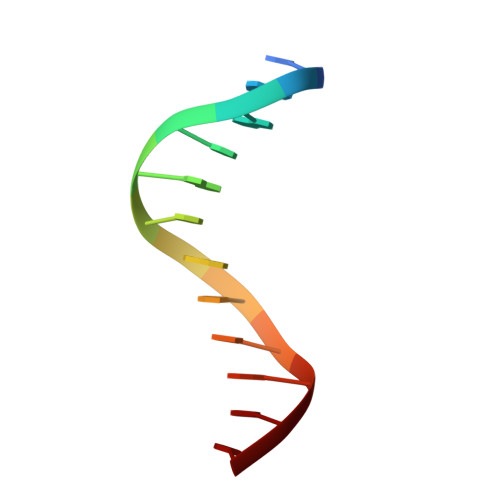
Structural basis of DNA target recognition by the B3 domain of Arabidopsis epigenome reader VAL1.
Sasnauskas, G., Kauneckaite, K., Siksnys, V.(2018) Nucleic Acids Res 46: 4316-4324
- PubMed: 29660015 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gky256
- Primary Citation Related Structures:
6FAS - PubMed Abstract:
Arabidopsis thaliana requires a prolonged period of cold exposure during winter to initiate flowering in a process termed vernalization. Exposure to cold induces epigenetic silencing of the FLOWERING LOCUS C (FLC) gene by Polycomb group (PcG) proteins. A key role in this epigenetic switch is played by transcriptional repressors VAL1 and VAL2, which specifically recognize Sph/RY DNA sequences within FLC via B3 DNA binding domains, and mediate recruitment of PcG silencing machinery. To understand the structural mechanism of site-specific DNA recognition by VAL1, we have solved the crystal structure of VAL1 B3 domain (VAL1-B3) bound to a 12 bp oligoduplex containing the canonical Sph/RY DNA sequence 5'-CATGCA-3'/5'-TGCATG-3'. We find that VAL1-B3 makes H-bonds and van der Waals contacts to DNA bases of all six positions of the canonical Sph/RY element. In agreement with the structure, in vitro DNA binding studies show that VAL1-B3 does not tolerate substitutions at any position of the 5'-TGCATG-3' sequence. The VAL1-B3-DNA structure presented here provides a structural model for understanding the specificity of plant B3 domains interacting with the Sph/RY and other DNA sequences.
- Institute of Biotechnology, Vilnius University, Sauletekio al. 7, LT-10257 Vilnius, Lithuania.
Organizational Affiliation: