Crystal structure of mammalian Rev7 in complex with human Rev3
Huber, F., Tropia, L., Emamzadah, S., Halazonetis, T.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mitotic spindle assembly checkpoint protein MAD2B | 211 | Mus musculus | Mutation(s): 5 Gene Names: Mad2l2, Mad2b, Rev7 |  | |
UniProt & NIH Common Fund Data Resources | |||||
IMPC: MGI:1919140 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9D752 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA polymerase zeta catalytic subunit | 28 | Homo sapiens | Mutation(s): 0 Gene Names: REV3L, POLZ, REV3 EC: 2.7.7.7 |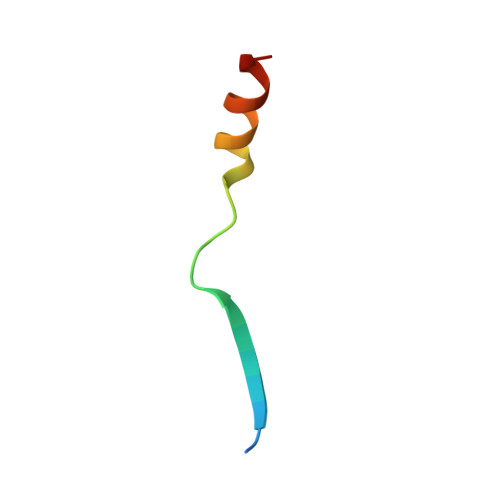  | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O60673 GTEx: ENSG00000009413 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O60673 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 39.2 | α = 90 |
| b = 48.64 | β = 90 |
| c = 116.81 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| iMOSFLM | data reduction |
| SCALA | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Swiss National Science Foundation | Switzerland | 160322 |