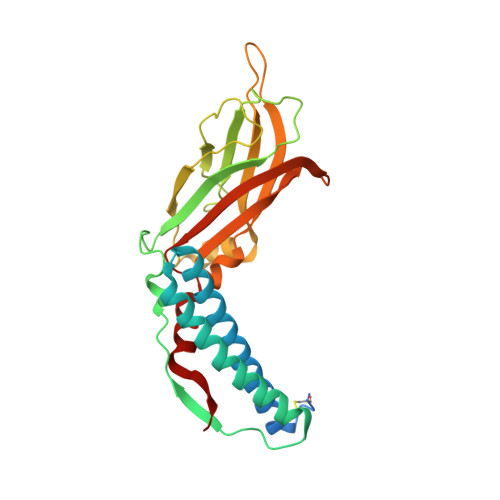
CC2D1B Coordinates ESCRT-III Activity during the Mitotic Reformation of the Nuclear Envelope.
Ventimiglia, L.N., Cuesta-Geijo, M.A., Martinelli, N., Caballe, A., Macheboeuf, P., Miguet, N., Parnham, I.M., Olmos, Y., Carlton, J.G., Weissenhorn, W., Martin-Serrano, J.(2018) Dev Cell 47: 547-563.e6
- PubMed: 30513301 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.devcel.2018.11.012
- Primary Citation Related Structures:
6EI6 - PubMed Abstract:
The coordinated reformation of the nuclear envelope (NE) after mitosis re-establishes the structural integrity and the functionality of the nuclear compartment. The endosomal sorting complex required for transport (ESCRT) machinery, a membrane remodeling pathway that is highly conserved in eukaryotes, has been recently involved in NE resealing by mediating the annular fusion of the nuclear membrane (NM). We show here that CC2D1B, a regulator of ESCRT polymerization, is required to re-establish the nuclear compartmentalization by coordinating endoplasmic reticulum (ER) membrane deposition around chromatin disks with ESCRT-III recruitment to the reforming NE. Accordingly, CC2D1B determines the spatiotemporal distribution of the CHMP7-ESCRT-III axis during NE reformation. Crucially, in CC2D1B-depleted cells, ESCRT activity is uncoupled from Spastin-mediated severing of spindle microtubules, resulting in persisting microtubules that compromise nuclear morphology. Therefore, we reveal CC2D1B as an essential regulatory factor that licenses the formation of ESCRT-III polymers to ensure the orderly reformation of the NE.
- Department of Infectious Diseases, King's College London, Faculty of Life Sciences & Medicine, London SE1 9RT, UK.
Organizational Affiliation: