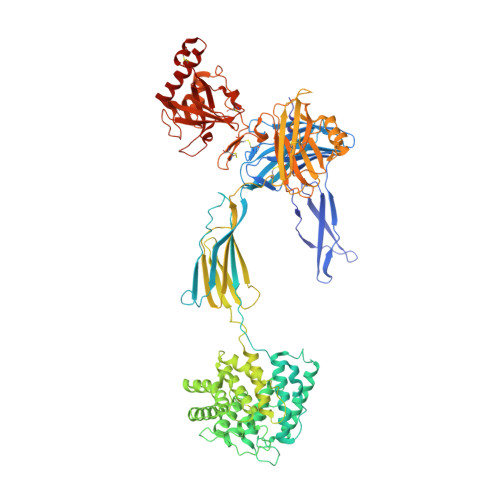
A potent complement factor C3-specific nanobody inhibiting multiple functions in the alternative pathway of human and murine complement.
Jensen, R.K., Pihl, R., Gadeberg, T.A.F., Jensen, J.K., Andersen, K.R., Thiel, S., Laursen, N.S., Andersen, G.R.(2018) J Biological Chem 293: 6269-6281
- PubMed: 29497000 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA117.001179
- Primary Citation Related Structures:
6EHG - PubMed Abstract:
The complement system is a complex, carefully regulated proteolytic cascade for which suppression of aberrant activation is of increasing clinical relevance, and inhibition of the complement alternative pathway is a subject of intense research. Here, we describe the nanobody hC3Nb1 that binds to multiple functional states of C3 with subnanomolar affinity. The nanobody causes a complete shutdown of alternative pathway activity in human and murine serum when present in concentrations comparable with that of C3, and hC3Nb1 is shown to prevent proconvertase assembly, as well as binding of the C3 substrate to C3 convertases. Our crystal structure of the C3b-hC3Nb1 complex and functional experiments demonstrate that proconvertase formation is blocked by steric hindrance between the nanobody and an Asn-linked glycan on complement factor B. In addition, hC3Nb1 is shown to prevent factor H binding to C3b, rationalizing its inhibition of factor I activity. Our results identify hC3Nb1 as a versatile, inexpensive, and powerful inhibitor of the alternative pathway in both human and murine in vitro model systems of complement activation.
- From the Departments of Molecular Biology and Genetics and.
Organizational Affiliation: