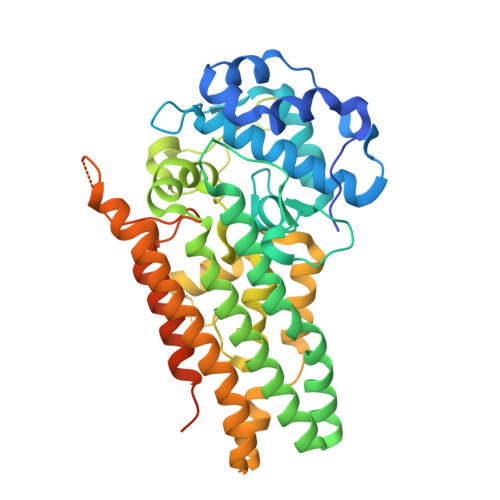
Crystal structure of human indoleamime 2,3-dioxygenase (IDO1) in complex with L-Trp and cyanide, Northeast Structural Genomics Target HR6160
Forouhar, F., Lewis-Ballester, A., Lew, S., Karkashon, S., Seetharaman, J., Lu, C., Hussain, M., Yeh, S.-R., Tong, L.To be published.