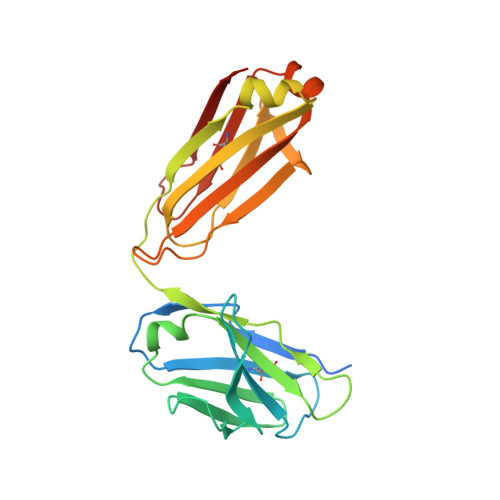
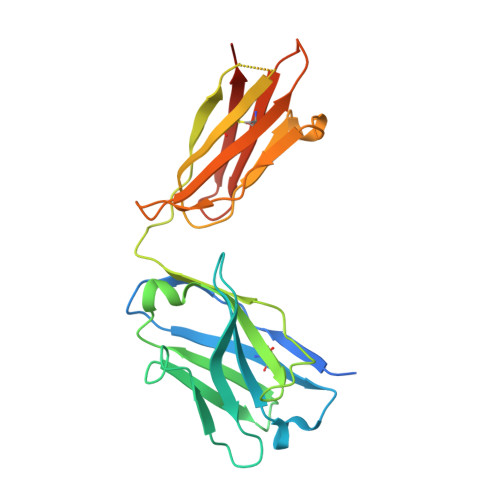
Structural investigation of human S. aureus-targeting antibodies that bind wall teichoic acid.
Fong, R., Kajihara, K., Chen, M., Hotzel, I., Mariathasan, S., Hazenbos, W.L.W., Lupardus, P.J.(2018) MAbs 10: 979-991
- PubMed: 30102105 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/19420862.2018.1501252
- Primary Citation Related Structures:
6DW2, 6DWA, 6DWC, 6DWI - PubMed Abstract:
Infections caused by methicillin-resistant Staphylococcus aureus (MRSA) are a growing health threat worldwide. Efforts to identify novel antibodies that target S. aureus cell surface antigens are a promising direction in the development of antibiotics that can halt MRSA infection. We biochemically and structurally characterized three patient-derived MRSA-targeting antibodies that bind to wall teichoic acid (WTA), which is a polyanionic surface glycopolymer. In S. aureus, WTA exists in both α- and β-forms, based on the stereochemistry of attachment of a N-acetylglucosamine residue to the repeating phosphoribitol sugar unit. We identified a panel of antibodies cloned from human patients that specifically recognize the α or β form of WTA, and can bind with high affinity to pathogenic wild-type strains of S. aureus bacteria. To investigate how the β-WTA specific antibodies interact with their target epitope, we determined the X-ray crystal structures of the three β-WTA specific antibodies, 4462, 4497, and 6078 (Protein Data Bank IDs 6DWI, 6DWA, and 6DW2, respectively), bound to a synthetic WTA epitope. These structures reveal that all three of these antibodies, while utilizing distinct antibody complementarity-determining region sequences and conformations to interact with β-WTA, fulfill two recognition principles: binding to the β-GlcNAc pyranose core and triangulation of WTA phosphate residues with polar contacts. These studies reveal the molecular basis for targeting a unique S. aureus cell surface epitope and highlight the power of human patient-based antibody discovery techniques for finding novel pathogen-targeting therapeutics.
- a Department of Structural Biology , Genentech , South San Francisco , CA , USA.
Organizational Affiliation: