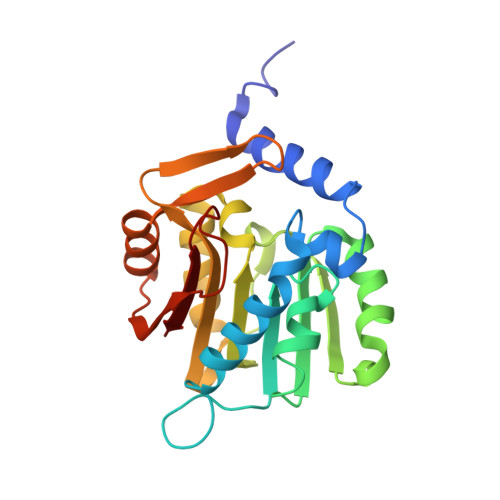
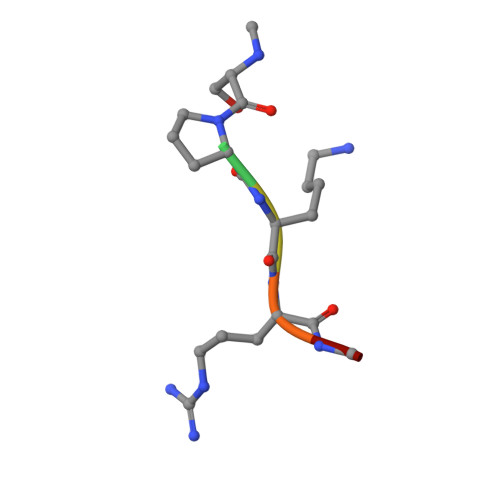
An asparagine/glycine switch governs product specificity of human N-terminal methyltransferase NTMT2.
Dong, C., Dong, G., Li, L., Zhu, L., Tempel, W., Liu, Y., Huang, R., Min, J.(2018) Commun Biol 1: 183-183
- PubMed: 30417120 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-018-0196-2
- Primary Citation Related Structures:
5UBB, 6DUB - PubMed Abstract:
α-N-terminal methylation of proteins is an important post-translational modification that is catalyzed by two different N-terminal methyltransferases, namely NTMT1 and NTMT2. Previous studies have suggested that NTMT1 is a tri-methyltransferase, whereas NTMT2 is a mono-methyltransferase. Here, we report the first crystal structures, to our knowledge, of NTMT2 in binary complex with S-adenosyl-L-methionine as well as in ternary complex with S-adenosyl-L-homocysteine and a substrate peptide. Our structural observations combined with biochemical studies reveal that NTMT2 is also able to di-/tri-methylate the GPKRIA peptide and di-methylate the PPKRIA peptide, otherwise it is predominantly a mono-methyltransferase. The residue N89 of NTMT2 serves as a gatekeeper residue that regulates the binding of unmethylated versus monomethylated substrate peptide. Structural comparison of NTMT1 and NTMT2 prompts us to design a N89G mutant of NTMT2 that can profoundly alter its catalytic activities and product specificities.
- Structural Genomics Consortium, University of Toronto, Toronto, M5G1L7, ON, Canada.
Organizational Affiliation: