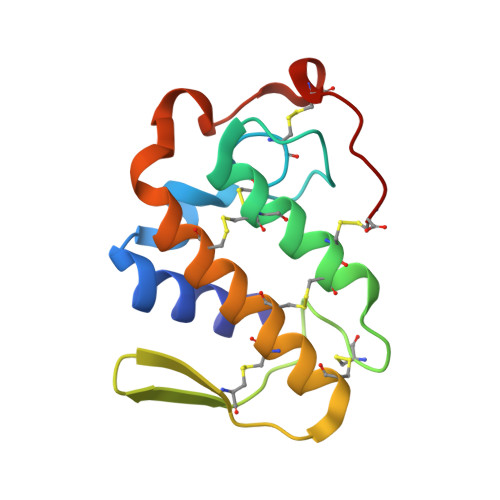
Structural basis of phospholipase A2-like myotoxin inhibition by chicoric acid, a novel potent inhibitor of ophidian toxins.
Cardoso, F.F., Borges, R.J., Dreyer, T.R., Salvador, G.H.M., Cavalcante, W.L.G., Pai, M.D., Gallacci, M., Fontes, M.R.M.(2018) Biochim Biophys Acta Gen Subj 1862: 2728-2737
- PubMed: 30251662 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbagen.2018.08.002
- Primary Citation Related Structures:
6DIK - PubMed Abstract:
Specific compounds found in vegetal species have been demonstrated to be efficient inhibitors of snake toxins, such as phospholipase A 2 -like (PLA 2 -like) proteins. These particular proteins, present in several species of vipers (Viperidae), induce a severe local myotoxic effect in prey and human victims, and this effect is often not efficiently neutralized by the regular serum therapy. PLA 2 -like proteins have been functionally and structurally studied since the early 1990s; however, a comprehensive molecular mechanism was proposed only recently. Myographic and histological techniques were used to evaluate the inhibitory effect of chicoric acid (CA) against BthTX-I myotoxin. Isothermal titration calorimetry assays were used to measure the affinity between the inhibitor and the toxin. X-ray crystallography was used to reveal details of this interaction. CA prevented the blockade of indirectly evoked muscle contraction and inhibited muscle damage induced by BthTX-I. The inhibitor binds to the toxin with the highest affinity measured for a natural compound in calorimetric assays. The crystal structure and molecular dynamics simulations demonstrated that CA binds at the entrance of the hydrophobic channel of the toxin and binds to one of the clusters that participates in membrane disruption. CA prevents the myotoxic activity of the toxin, preventing its activation by simultaneous binding with two critical regions. CA is a potential myotoxic inhibitor to other PLA 2 -like proteins and a possible candidate to complement serum therapy.
- Departamento de Física e Biofísica, Instituto de Biociências, Universidade Estadual Paulista (UNESP), Botucatu, SP, Brazil.
Organizational Affiliation: