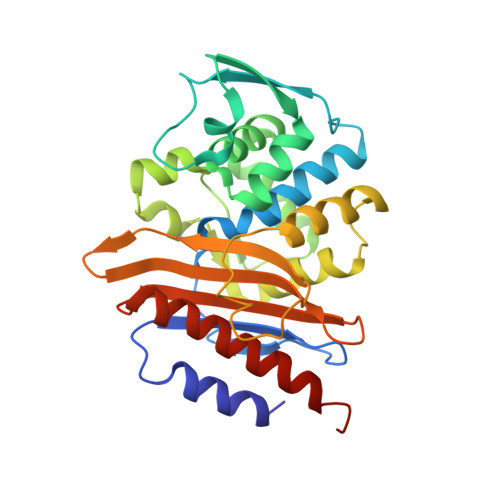
Structural Insights into the Inhibition of the Extended-Spectrum beta-Lactamase PER-2 by Avibactam.
Ruggiero, M., Papp-Wallace, K.M., Brunetti, F., Barnes, M.D., Bonomo, R.A., Gutkind, G., Klinke, S., Power, P.(2019) Antimicrob Agents Chemother 63
- PubMed: 31235626 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/AAC.00487-19
- Primary Citation Related Structures:
6D3G, 6DGU - PubMed Abstract:
The diazabicyclooctane (DBO) avibactam (AVI) reversibly inactivates most serine-β-lactamases. Previous investigations showed that inhibition constants of AVI toward class A PER-2 are reminiscent of values observed for class C and D β-lactamases (i.e., k 2 / K of ≈10 3 M -1 s -1 ) but lower than other class A β-lactamases (i.e., k 2 / K = 10 4 to 10 5 M -1 s -1 ). Herein, biochemical and structural studies were conducted with PER-2 and AVI to explore these differences. Furthermore, biochemical studies on Arg220 and Thr237 variants with AVI were conducted to gain deeper insight into the mechanism of PER-2 inactivation. The main biochemical and structural observations revealed the following: (i) both amino-acid substitutions in Arg220 and the rich hydrophobic content in the active site hinder the binding of catalytic waters and acylation, impairing AVI inhibition; (ii) movement of Ser130 upon binding of AVI favors the formation of a hydrogen bond with the sulfate group of AVI; and (iii) the Thr237Ala substitution alters the AVI inhibition constants. The acylation constant ( k 2 / K ) of PER-2 by AVI is primarily influenced by stabilizing hydrogen bonds involving AVI and important residues such as Thr237 and Arg220. (Variants in Arg220 demonstrate a dramatic reduction in k 2 / K ) We also observed that displacement of Ser130 side chain impairs AVI acylation, an observation not made in other extended-spectrum β-lactamases (ESBLs). Comparatively, relebactam combined with a β-lactam is more potent against Escherichia coli producing PER-2 variants than β-lactam-AVI combinations. Our findings provide a rationale for evaluating the utility of the currently available DBO inhibitors against unique ESBLs like PER-2 and anticipate the effectiveness of these inhibitors toward variants that may eventually be selected upon AVI usage.
- Universidad de Buenos Aires, Facultad de Farmacia y Bioquímica, Departamento de Microbiología, Inmunología y Biotecnología, Laboratorio de Resistencia Bacteriana, Buenos Aires, Argentina.
Organizational Affiliation: