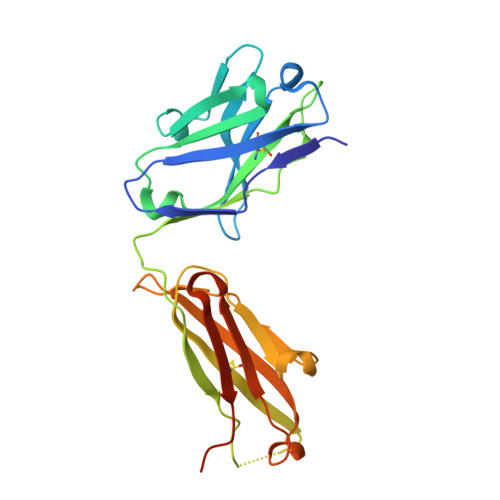
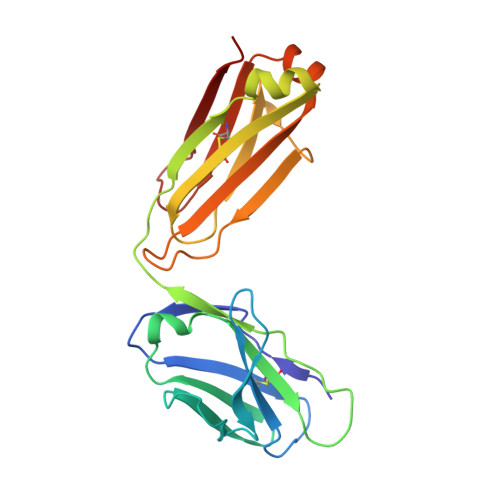
Structural Characterization of Pan-Ebolavirus Antibody 6D6 Targeting the Fusion Peptide of the Surface Glycoprotein.
Milligan, J.C., Parekh, D.V., Fuller, K.M., Igarashi, M., Takada, A., Saphire, E.O.(2019) J Infect Dis 219: 415-419
- PubMed: 30203042 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/infdis/jiy532
- Primary Citation Related Structures:
6DG2 - PubMed Abstract:
Ebola virus infection causes severe disease in humans and represents a global health threat. Candidates for immunotherapeutics and vaccines have shown promise in clinical trials, although they are ineffective against other members of the Ebolavirus genus that also cause periodic, lethal outbreaks. In this study, we present a crystal structure of a pan-ebolavirus antibody, 6D6, as well as single-particle electron microscopy reconstructions of 6D6 in complex with Ebola and Bundibugyo virus glycoproteins. 6D6 binds to the conserved glycoprotein fusion peptide, implicating it as a site of immune vulnerability that could be exploited to reliably elicit a pan-ebolavirus neutralizing antibody response.
- Department of Immunology and Microbiology, La Jolla, California.
Organizational Affiliation: