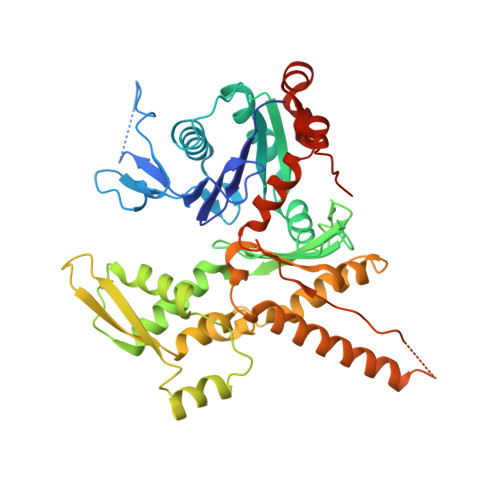
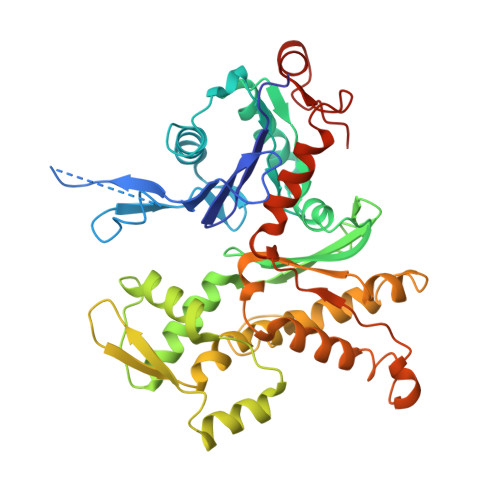
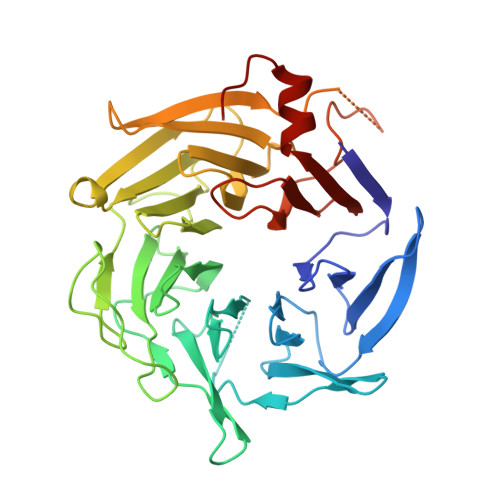
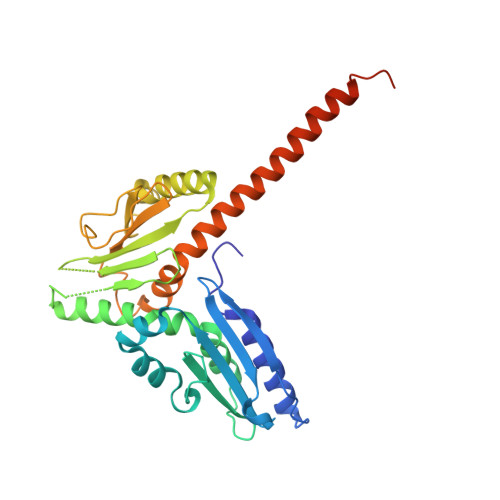
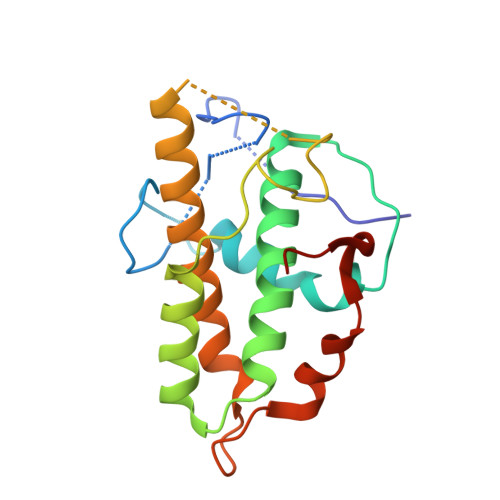
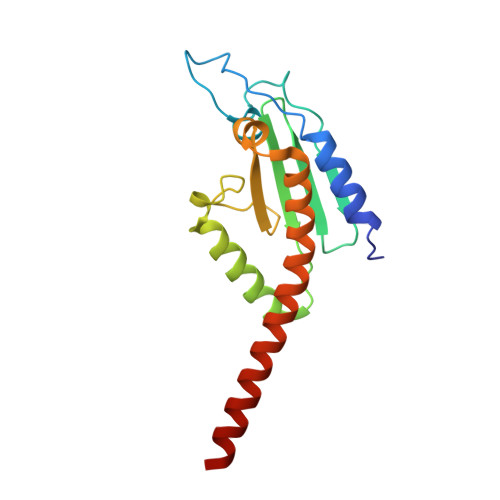
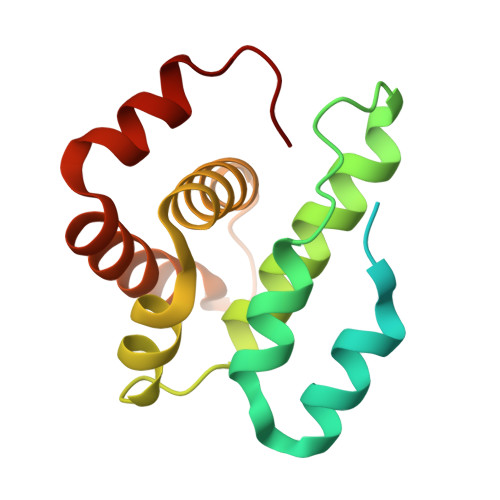
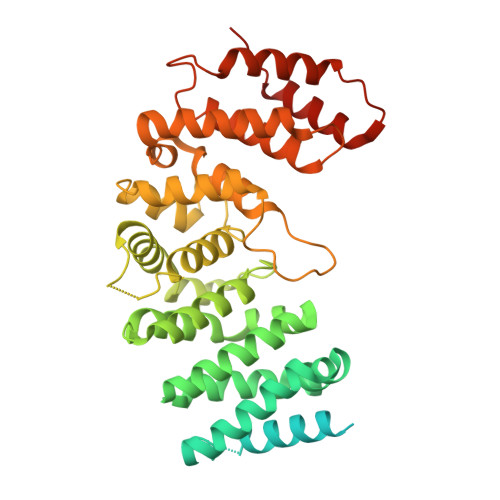
Structure of the nucleation-promoting factor SPIN90 bound to the actin filament nucleator Arp2/3 complex.
Luan, Q., Liu, S.L., Helgeson, L.A., Nolen, B.J.(2018) EMBO J 37
- PubMed: 30322896 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2018100005
- Primary Citation Related Structures:
6DEC, 6DED, 6DEE - PubMed Abstract:
Unlike the WASP family of Arp2/3 complex activators, WISH/DIP/SPIN90 (WDS) family proteins activate actin filament nucleation by the Arp2/3 complex without the need for a preformed actin filament. This allows WDS proteins to initiate branched actin network assembly by providing seed filaments that activate WASP-bound Arp2/3 complex. Despite their important role in actin network initiation, it is unclear how WDS proteins drive the activating steps that require both WASP and pre-existing actin filaments during WASP-mediated nucleation. Here, we show that SPIN90 folds into an armadillo repeat domain that binds a surface of Arp2/3 complex distinct from the two WASP sites, straddling a hinge point that may stimulate movement of the Arp2 subunit into the activated short-pitch conformation. SPIN90 binds a surface on Arp2/3 complex that overlaps with actin filament binding, explaining how it could stimulate the same structural rearrangements in the complex as pre-existing actin filaments. By revealing how WDS proteins activate the Arp2/3 complex, these data provide a molecular foundation to understand initiation of dendritic actin networks and regulation of Arp2/3 complex by its activators.
- Institute of Molecular Biology and Department of Chemistry and Biochemistry, University of Oregon, Eugene, OR, USA.
Organizational Affiliation: