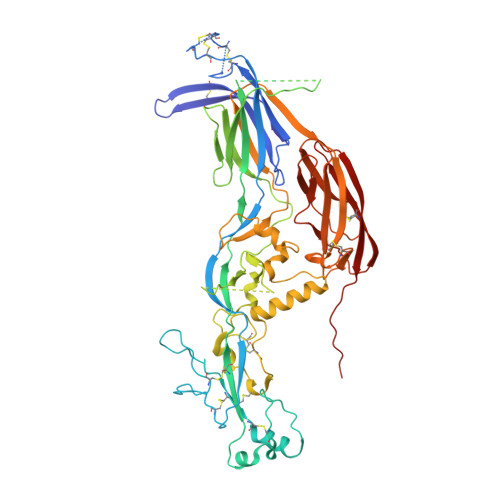
Fusion surface structure, function, and dynamics of gamete fusogen HAP2.
Feng, J., Dong, X., Pinello, J., Zhang, J., Lu, C., Iacob, R.E., Engen, J.R., Snell, W.J., Springer, T.A.(2018) Elife 7
- PubMed: 30281023 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.39772
- Primary Citation Related Structures:
6DBS - PubMed Abstract:
HAP2 is a class II gamete fusogen in many eukaryotic kingdoms. A crystal structure of Chlamydomonas HAP2 shows a trimeric fusion state. Domains D1, D2.1 and D2.2 line the 3-fold axis; D3 and a stem pack against the outer surface. Surprisingly, hydrogen-deuterium exchange shows that surfaces of D1, D2.2 and D3 closest to the 3-fold axis are more dynamic than exposed surfaces. Three fusion helices in the fusion loops of each monomer expose hydrophobic residues at the trimer apex that are splayed from the 3-fold axis, leaving a solvent-filled cavity between the fusion loops in each monomer. At the base of the two fusion loops, Arg185 docks in a carbonyl cage. Comparisons to other structures, dynamics, and the greater effect on Chlamydomonas gamete fusion of mutation of axis-proximal than axis-distal fusion helices suggest that the apical portion of each monomer could tilt toward the 3-fold axis with merger of the fusion helices into a common fusion surface.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, United States.
Organizational Affiliation: