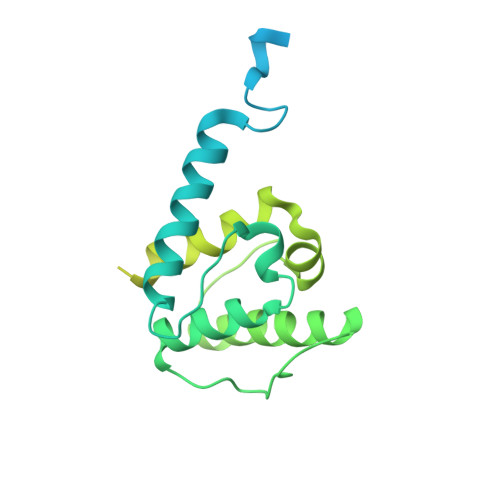
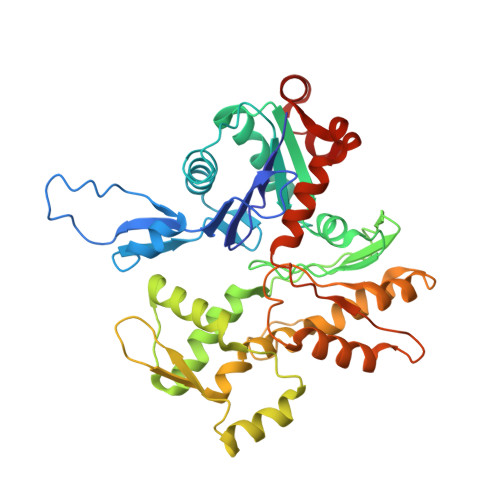
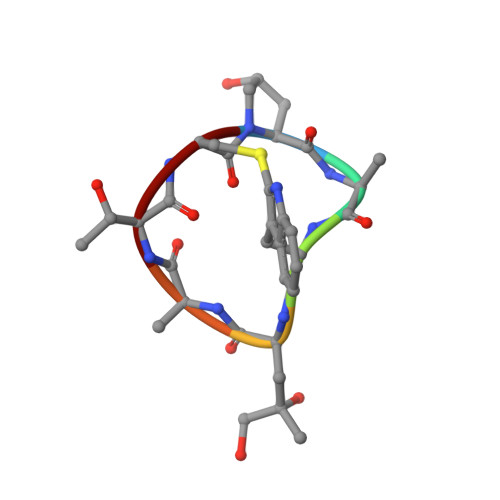
Structural basis of the filamin A actin-binding domain interaction with F-actin.
Iwamoto, D.V., Huehn, A., Simon, B., Huet-Calderwood, C., Baldassarre, M., Sindelar, C.V., Calderwood, D.A.(2018) Nat Struct Mol Biol 25: 918-927
- PubMed: 30224736 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-018-0128-3
- Primary Citation Related Structures:
6D8C - PubMed Abstract:
Actin-cross-linking proteins assemble actin filaments into higher-order structures essential for orchestrating cell shape, adhesion, and motility. Missense mutations in the tandem calponin homology domains of their actin-binding domains (ABDs) underlie numerous genetic diseases, but a molecular understanding of these pathologies is hampered by the lack of high-resolution structures of any actin-cross-linking protein bound to F-actin. Here, taking advantage of a high-affinity, disease-associated mutant of the human filamin A (FLNa) ABD, we combine cryo-electron microscopy and functional studies to reveal at near-atomic resolution how the first calponin homology domain (CH1) and residues immediately N-terminal to it engage actin. We further show that reorientation of CH2 relative to CH1 is required to avoid clashes with actin and to expose F-actin-binding residues on CH1. Our data explain localization of disease-associated loss-of-function mutations to FLNaCH1 and gain-of-function mutations to the regulatory FLNaCH2. Sequence conservation argues that this provides a general model for ABD-F-actin binding.
- Department of Pharmacology, Yale University, New Haven, CT, USA.
Organizational Affiliation: