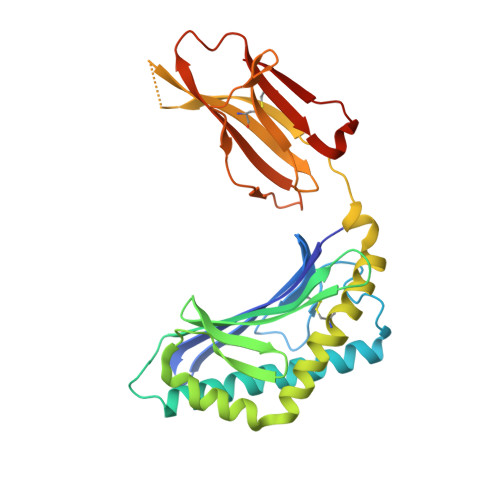
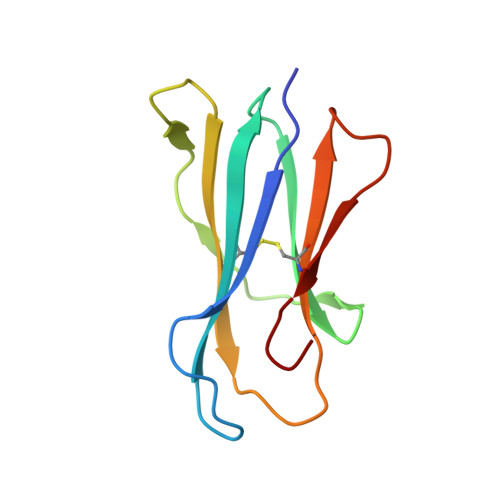
Control of CD1d-restricted antigen presentation and inflammation by sphingomyelin.
Melum, E., Jiang, X., Baker, K.D., Macedo, M.F., Fritsch, J., Dowds, C.M., Wang, J., Pharo, A., Kaser, A., Tan, C., Pereira, C.S., Kelly, S.L., Duan, J., Karlsen, T.H., Exley, M.A., Schutze, S., Zajonc, D.M., Merrill, A.H., Schuchman, E.H., Zeissig, S., Blumberg, R.S.(2019) Nat Immunol 20: 1644-1655
- PubMed: 31636468 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41590-019-0504-0
- Primary Citation Related Structures:
6CYW - PubMed Abstract:
Invariant natural killer T (iNKT) cells recognize activating self and microbial lipids presented by CD1d. CD1d can also bind non-activating lipids, such as sphingomyelin. We hypothesized that these serve as endogenous regulators and investigated humans and mice deficient in acid sphingomyelinase (ASM), an enzyme that degrades sphingomyelin. We show that ASM absence in mice leads to diminished CD1d-restricted antigen presentation and iNKT cell selection in the thymus, resulting in decreased iNKT cell levels and resistance to iNKT cell-mediated inflammatory conditions. Defective antigen presentation and decreased iNKT cells are also observed in ASM-deficient humans with Niemann-Pick disease, and ASM activity in healthy humans correlates with iNKT cell phenotype. Pharmacological ASM administration facilitates antigen presentation and restores the levels of iNKT cells in ASM-deficient mice. Together, these results demonstrate that control of non-agonistic CD1d-associated lipids is critical for iNKT cell development and function in vivo and represents a tight link between cellular sphingolipid metabolism and immunity.
- Gastroenterology Division, Department of Medicine, Brigham and Women's Hospital, Harvard Medical School, Boston, MA, USA. espen.melum@medisin.uio.no.
Organizational Affiliation: