Crystal structure of Signal recognition particle receptor FtsY from Elizabethkingia anophelis in complex with GDP
Delker, S.L., Abendroth, J., Lorimer, D., Edwards, T.E.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Signal recognition particle receptor FtsY | 329 | Elizabethkingia anophelis NUHP1 | Mutation(s): 0 Gene Names: ftsY, BD94_0405 EC: 3.6.5.4 | 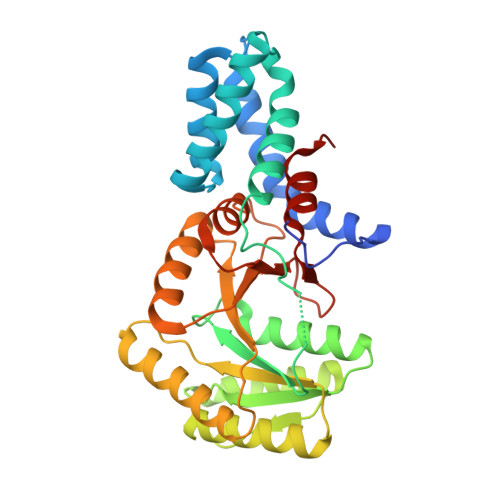 | |
UniProt | |||||
Find proteins for A0A077EFA5 (Elizabethkingia anophelis NUHP1) Explore A0A077EFA5 Go to UniProtKB: A0A077EFA5 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A077EFA5 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GDP (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth A], E [auth A], J [auth B] | GUANOSINE-5'-DIPHOSPHATE C10 H15 N5 O11 P2 QGWNDRXFNXRZMB-UUOKFMHZSA-N |  | ||
| EDO Download:Ideal Coordinates CCD File | F [auth A] G [auth A] H [auth A] I [auth A] L [auth B] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| MG Download:Ideal Coordinates CCD File | D [auth A], K [auth B] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 49.52 | α = 101.18 |
| b = 49.64 | β = 99.1 |
| c = 69.71 | γ = 105.58 |
| Software Name | Purpose |
|---|---|
| XSCALE | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| MOLREP | phasing |