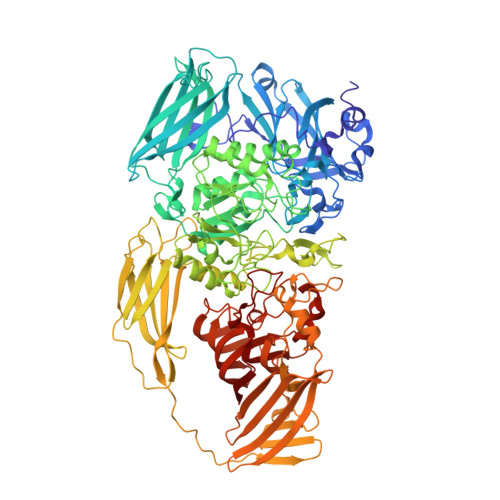
Atomic Resolution Cryo-EM Structure of beta-Galactosidase.
Bartesaghi, A., Aguerrebere, C., Falconieri, V., Banerjee, S., Earl, L.A., Zhu, X., Grigorieff, N., Milne, J.L.S., Sapiro, G., Wu, X., Subramaniam, S.(2018) Structure 26: 848
- PubMed: 29754826 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2018.04.004
- Primary Citation Related Structures:
6CVM - PubMed Abstract:
The advent of direct electron detectors has enabled the routine use of single-particle cryo-electron microscopy (EM) approaches to determine structures of a variety of protein complexes at near-atomic resolution. Here, we report the development of methods to account for local variations in defocus and beam-induced drift, and the implementation of a data-driven dose compensation scheme that significantly improves the extraction of high-resolution information recorded during exposure of the specimen to the electron beam. These advances enable determination of a cryo-EM density map for β-galactosidase bound to the inhibitor phenylethyl β-D-thiogalactopyranoside where the ordered regions are resolved at a level of detail seen in X-ray maps at ∼ 1.5 Å resolution. Using this density map in conjunction with constrained molecular dynamics simulations provides a measure of the local flexibility of the non-covalently bound inhibitor and offers further opportunities for structure-guided inhibitor design.
- Laboratory of Cell Biology, Center for Cancer Research, National Cancer Institute, NIH, Bethesda, MD 20892, USA.
Organizational Affiliation: