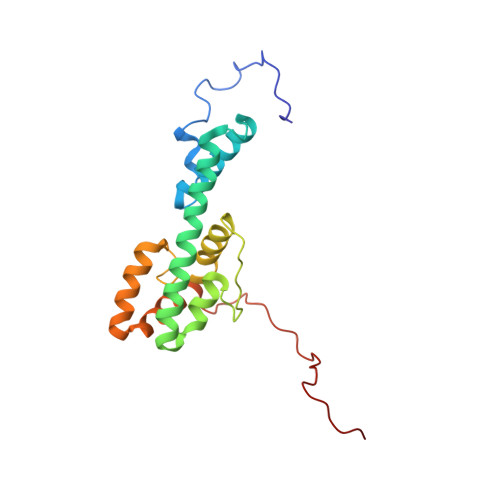
The Structure of Melanoregulin Reveals a Role for Cholesterol Recognition in the Protein's Ability to Promote Dynein Function.
Rout, A.K., Wu, X., Starich, M.R., Strub, M.P., Hammer, J.A., Tjandra, N.(2018) Structure 26: 1373
- PubMed: 30174147 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2018.07.009
- Primary Citation Related Structures:
6CMY - PubMed Abstract:
Melanoregulin (Mreg) is a small, highly charged, multiply palmitoylated protein present on the membrane of melanosomes. Mreg is implicated in the transfer of melanosomes from melanocytes to keratinocytes, and in promoting the microtubule minus end-directed transport of these organelles. The possible molecular function of Mreg was identified by solving its structure using nuclear magnetic resonance (NMR) spectroscopy. Mreg contains six α helices forming a fishhook-like fold in which positive and negative charges occupy opposite sides of the protein's surface and sandwich a putative, cholesterol recognition sequence (CRAC motif). Mreg containing a point mutation within its CRAC motif still targets to late endosomes/lysosomes, but no longer promotes their microtubule minus end-directed transport. Moreover, wild-type Mreg does not promote the microtubule minus end-directed transport of late endosomes/lysosomes in cells transiently depleted of cholesterol. Finally, reversing the charge of three clustered acidic residues partially inhibits Mreg's ability to drive these organelles to microtubule minus ends.
- Biochemistry and Biophysics Center, National Heart, Lung, and Blood Institute, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: