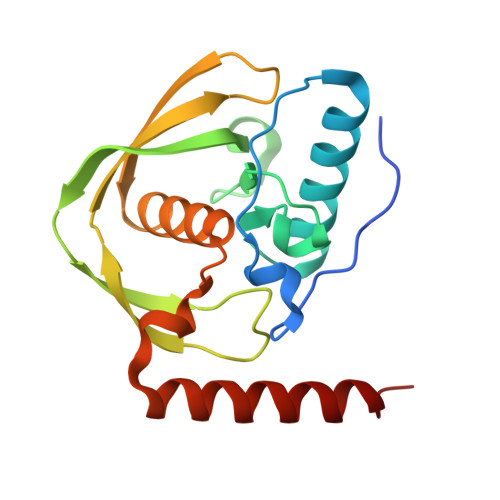
Structures of Legionella pneumophila serogroup 1 peptide deformylase bound to nickel(II) and actinonin.
Nguyen, C.L., Fan, W., Fisher, S., Matthews, K., Norman, J.O., Abendroth, J., Barrett, K.F., Craig, J.K., Edwards, T.E., Lorimer, D.D., McLaughlin, K.J.(2025) Acta Crystallogr F Struct Biol Commun 81: 163-170
- PubMed: 40091854 Search on PubMed
- DOI: https://doi.org/10.1107/S2053230X25001876
- Primary Citation Related Structures:
6CK7 - PubMed Abstract:
Legionella pneumophila serogroup 1 is the primary causative agent of Legionnaires' disease, a rare but severe respiratory infection. While the fatality rate of Legionnaires' disease is low in the general population, it is more pronounced in vulnerable communities such as the immunocompromised. Thus, the development of new antimicrobials is of interest for use when existing antibiotics may not be applicable. Peptide deformylases (PDFs) have been under continued investigation as targets for novel antimicrobial compounds. PDF plays an essential role in protein synthesis, removing the N-terminal formyl group from new polypeptides, and is required for growth in most bacteria. Here, we report two crystal structures of L. pneumophila serogroup 1 PDF (LpPDF) bound to either Ni 2+ , an active state, or inhibited by actinonin and Zn 2+ ; the structures were determined to 1.5 and 1.65 Å resolution, respectively, and were solved by the Seattle Structural Genomics Center for Infectious Disease (SSGCID). The SSGCID is charged with determining structures of biologically important proteins and molecules from human pathogens. As actinonin is an antimicrobial natural product that has been used as a reference compound in drug development, these structures will help support the ongoing drug-development process.
- Biochemistry Program, Vassar College, 124 Raymond Avenue, Poughkeepsie, NY 12604, USA.
Organizational Affiliation: