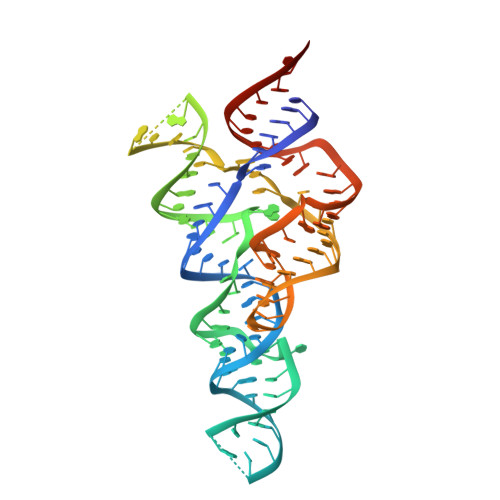
Structures of two aptamers with differing ligand specificity reveal ruggedness in the functional landscape of RNA.
Knappenberger, A.J., Reiss, C.W., Strobel, S.A.(2018) Elife 7
- PubMed: 29877798 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.36381
- Primary Citation Related Structures:
6CK4, 6CK5 - PubMed Abstract:
Two classes of riboswitches related to the ykkC guanidine-I riboswitch bind phosphoribosyl pyrophosphate (PRPP) and guanosine tetraphosphate (ppGpp). Here we report the co-crystal structure of the PRPP aptamer and its ligand. We also report the structure of the G96A point mutant that prefers ppGpp over PRPP with a dramatic 40,000-fold switch in specificity. The ends of the aptamer form a helix that is not present in the guanidine aptamer and is involved in the expression platform. In the mutant, the base of ppGpp replaces G96 in three-dimensional space. This disrupts the S-turn, which is a primary structural feature of the ykkC RNA motif. These dramatic differences in ligand specificity are achieved with minimal mutations. ykkC aptamers are therefore a prime example of an RNA fold with a rugged fitness landscape. The ease with which the ykkC aptamer acquires new specificity represents a striking case of evolvability in RNA.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, United States.
Organizational Affiliation: