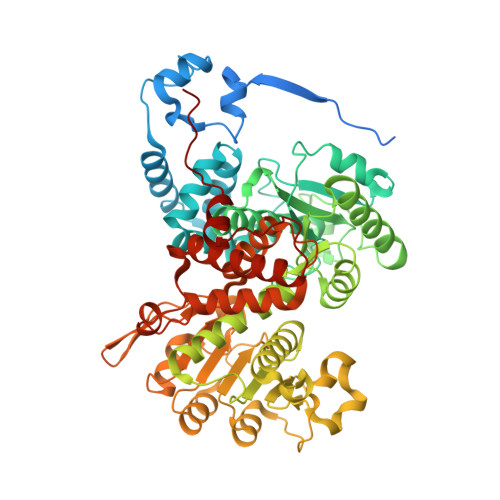
Molecular adaptations of NADP-malic enzyme for its function in C4photosynthesis in grasses.
Alvarez, C.E., Bovdilova, A., Hoppner, A., Wolff, C.C., Saigo, M., Trajtenberg, F., Zhang, T., Buschiazzo, A., Nagel-Steger, L., Drincovich, M.F., Lercher, M.J., Maurino, V.G.(2019) Nat Plants 5: 755-765
- PubMed: 31235877 Search on PubMed
- DOI: https://doi.org/10.1038/s41477-019-0451-7
- Primary Citation Related Structures:
5OU5, 6C7N - PubMed Abstract:
In C 4 grasses of agronomical interest, malate shuttled into the bundle sheath cells is decarboxylated mainly by nicotinamide adenine dinucleotide phosphate (NADP)-malic enzyme (C 4 -NADP-ME). The activity of C 4 -NADP-ME was optimized by natural selection to efficiently deliver CO 2 to Rubisco. During its evolution from a plastidic non-photosynthetic NADP-ME, C 4 -NADP-ME acquired increased catalytic efficiency, tetrameric structure and pH-dependent inhibition by its substrate malate. Here, we identified specific amino acids important for these C 4 adaptions based on strict differential conservation of amino acids, combined with solving the crystal structures of maize and sorghum C 4 -NADP-ME. Site-directed mutagenesis and structural analyses show that Q503, L544 and E339 are involved in catalytic efficiency; E339 confers pH-dependent regulation by malate, F140 is critical for the stabilization of the oligomeric structure and the N-terminal region is involved in tetramerization. Together, the identified molecular adaptations form the basis for the efficient catalysis and regulation of one of the central biochemical steps in C 4 metabolism.
- Centro de Estudios Fotosinteticos y Bioquimicos (CEFOBI-CONICET), Facultad de Ciencias Bioquímicas y Farmacéuticas, University of Rosario, Rosario, Argentina.
Organizational Affiliation: