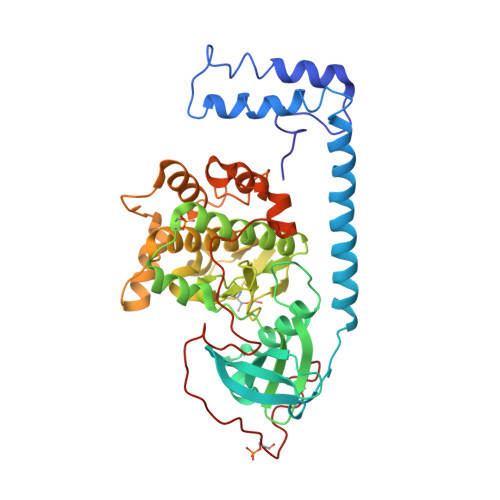
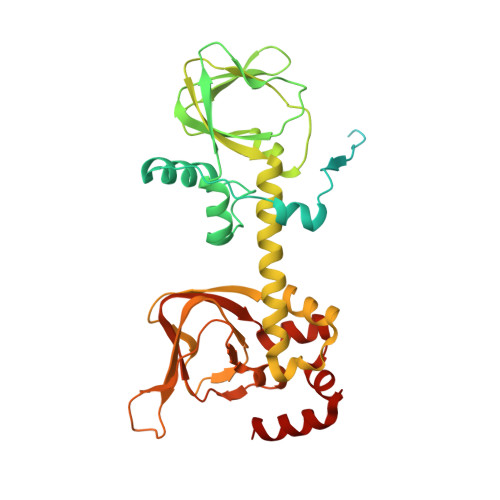
Structures of the PKA RI alpha Holoenzyme with the FLHCC Driver J-PKAc alpha or Wild-Type PKAc alpha.
Cao, B., Lu, T.W., Martinez Fiesco, J.A., Tomasini, M., Fan, L., Simon, S.M., Taylor, S.S., Zhang, P.(2019) Structure 27: 816
- PubMed: 30905674 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2019.03.001
- Primary Citation Related Structures:
6BYR, 6BYS - PubMed Abstract:
Fibrolamellar hepatocellular carcinoma (FLHCC) is driven by J-PKAcα, a kinase fusion chimera of the J domain of DnaJB1 with PKAcα, the catalytic subunit of protein kinase A (PKA). Here we report the crystal structures of the chimeric fusion RIα 2 :J-PKAcα 2 holoenzyme formed by J-PKAcα and the PKA regulatory (R) subunit RIα, and the wild-type (WT) RIα 2 :PKAcα 2 holoenzyme. The chimeric and WT RIα holoenzymes have quaternary structures different from the previously solved WT RIβ and RIIβ holoenzymes. The WT RIα holoenzyme showed the same configuration as the chimeric RIα 2 :J-PKAcα 2 holoenzyme and a distinct second conformation. The J domains are positioned away from the symmetrical interface between the two RIα:J-PKAcα heterodimers in the chimeric fusion holoenzyme and are highly dynamic. The structural and dynamic features of these holoenzymes enhance our understanding of the fusion chimera protein J-PKAcα that drives FLHCC as well as the isoform specificity of PKA.
- Structural Biophysics Laboratory, Center for Cancer Research, National Cancer Institute, Frederick, MD, USA.
Organizational Affiliation: