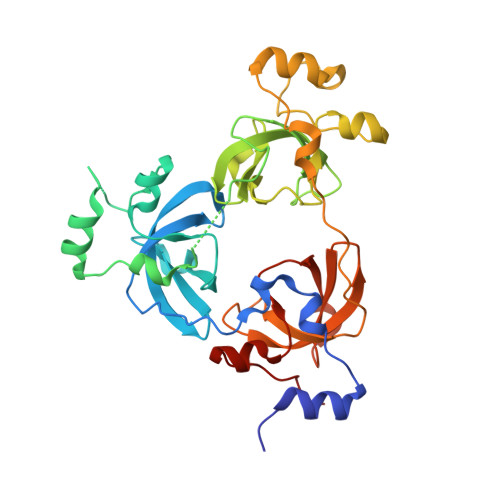
Crystal structure of L3MBTL1 MBT Domain with MBK14970
DOBROVETSKY, E., DONG, A., NICHOLSON, B., COX, C., FISCHER, C., ARMACOST, K., SANDERS, J., Bountra, C., Arrowsmith, C.H., Edwards, A.M., BROWN, P.J., Structural Genomics Consortium (SGC)To be published.