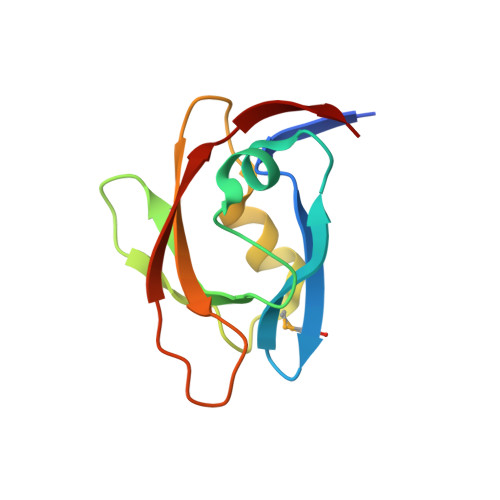
1.45 Angstrom Resolution Crystal Structure of PDZ domain of Carboxy-Terminal Protease from Vibrio cholerae in Complex with Peptide.
Minasov, G., Shuvalova, L., Filippova, E.V., Kiryukhina, O., Grimshaw, S., Kwon, K., Anderson, W.F., Satchell, K.J.F., Joachimiak, A., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.