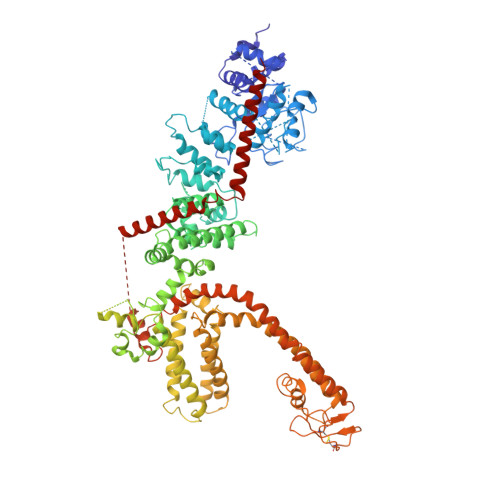
Structure of full-length human TRPM4.
Duan, J., Li, Z., Li, J., Santa-Cruz, A., Sanchez-Martinez, S., Zhang, J., Clapham, D.E.(2018) Proc Natl Acad Sci U S A 115: 2377-2382
- PubMed: 29463718 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1722038115
- Primary Citation Related Structures:
6BWI - PubMed Abstract:
Transient receptor potential melastatin subfamily member 4 (TRPM4) is a widely distributed, calcium-activated, monovalent-selective cation channel. Mutations in human TRPM4 (hTRPM4) result in progressive familial heart block. Here, we report the electron cryomicroscopy structure of hTRPM4 in a closed, Na + -bound, apo state at pH 7.5 to an overall resolution of 3.7 Å. Five partially hydrated sodium ions are proposed to occupy the center of the conduction pore and the entrance to the coiled-coil domain. We identify an upper gate in the selectivity filter and a lower gate at the entrance to the cytoplasmic coiled-coil domain. Intramolecular interactions exist between the TRP domain and the S4-S5 linker, N-terminal domain, and N and C termini. Finally, we identify aromatic interactions via π-π bonds and cation-π bonds, glycosylation at an N-linked extracellular site, a pore-loop disulfide bond, and 24 lipid binding sites. We compare and contrast this structure with other TRP channels and discuss potential mechanisms of regulation and gating of human full-length TRPM4.
- Howard Hughes Medical Institute, Ashburn, VA 20147.
Organizational Affiliation: