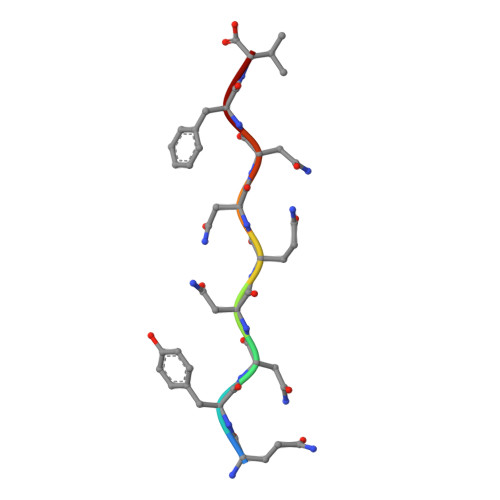
Sub-angstrom cryo-EM structure of a prion protofibril reveals a polar clasp.
Gallagher-Jones, M., Glynn, C., Boyer, D.R., Martynowycz, M.W., Hernandez, E., Miao, J., Zee, C.T., Novikova, I.V., Goldschmidt, L., McFarlane, H.T., Helguera, G.F., Evans, J.E., Sawaya, M.R., Cascio, D., Eisenberg, D.S., Gonen, T., Rodriguez, J.A.(2018) Nat Struct Mol Biol 25: 131-134
- PubMed: 29335561 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-017-0018-0
- Primary Citation Related Structures:
6AXZ, 6BTK - PubMed Abstract:
The atomic structure of the infectious, protease-resistant, β-sheet-rich and fibrillar mammalian prion remains unknown. Through the cryo-EM method MicroED, we reveal the sub-ångström-resolution structure of a protofibril formed by a wild-type segment from the β2-α2 loop of the bank vole prion protein. The structure of this protofibril reveals a stabilizing network of hydrogen bonds that link polar zippers within a sheet, producing motifs we have named 'polar clasps'.
- Department of Chemistry and Biochemistry, UCLA-DOE Institute for Genomics and Proteomics, University of California Los Angeles, Los Angeles, CA, USA.
Organizational Affiliation: