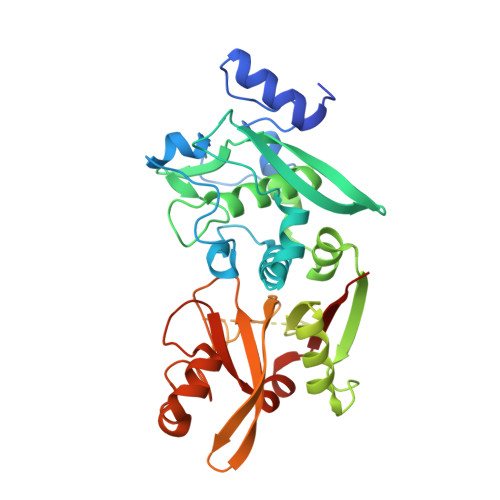
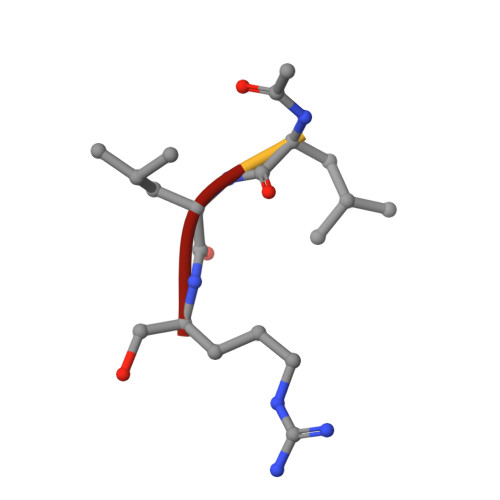
Structures of human calpain-3 protease core with and without bound inhibitor reveal mechanisms of calpain activation.
Ye, Q., Campbell, R.L., Davies, P.L.(2018) J Biological Chem 293: 4056-4070
- PubMed: 29382717 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA117.001097
- Primary Citation Related Structures:
6BDT, 6BGP, 6BJD, 6BKJ - PubMed Abstract:
Limb-girdle muscular dystrophy type 2a arises from mutations in the Ca 2+ -activated intracellular cysteine protease calpain-3. This calpain isoform is abundant in skeletal muscle and differs from the main isoforms, calpain-1 and -2, in being a homodimer and having two short insertion sequences. The first of these, IS1, interrupts the protease core and must be cleaved for activation and substrate binding. Here, to learn how calpain-3 can be regulated and inhibited, we determined the structures of the calpain-3 protease core with IS1 present or proteolytically excised. To prevent intramolecular IS1 autoproteolysis, we converted the active-site Cys to Ala. Small-angle X-ray scattering (SAXS) analysis of the C129A mutant suggested that IS1 is disordered and mobile enough to occupy several locations. Surprisingly, this was also true for the apo version of this mutant. We therefore concluded that IS1 might have a binding partner in the sarcomere and is unstructured in its absence. After autoproteolytic IS1 removal from the active Cys 129 calpain-3 protease core, we could solve its crystal structures with and without the cysteine protease inhibitors E-64 and leupeptin covalently bound to the active-site cysteine. In each structure, the active state of the protease core was assembled by the cooperative binding of two Ca 2+ ions to the equivalent sites used in calpain-1 and -2. These structures of the calpain-3 active site with residual IS1 and with bound E-64 and leupeptin may help guide the design of calpain-3-specific inhibitors.
- From the Department of Biomedical and Molecular Sciences, Queen's University, Kingston, Ontario K7L 3N6, Canada.
Organizational Affiliation: