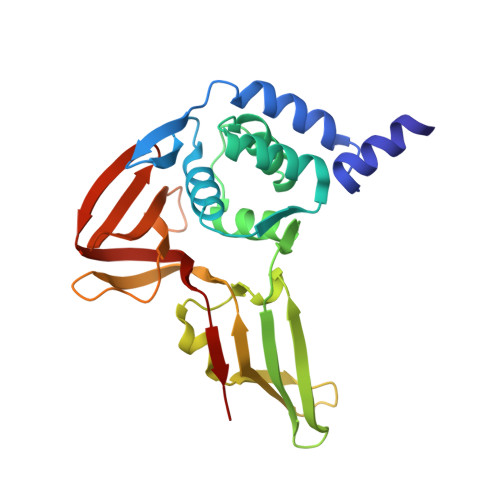
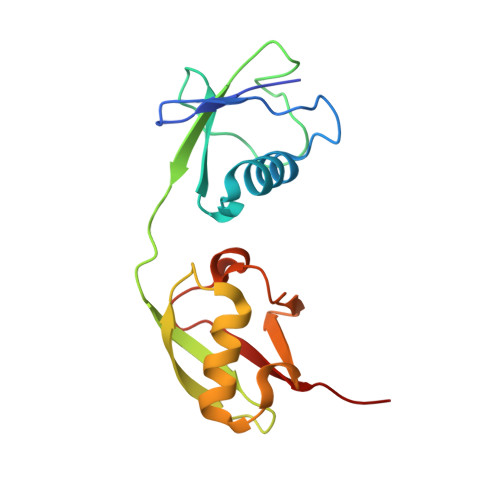
Decoupling deISGylating and deubiquitinating activities of the MERS virus papain-like protease.
Clasman, J.R., Everett, R.K., Srinivasan, K., Mesecar, A.D.(2020) Antiviral Res 174: 104661-104661
- PubMed: 31765674 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.antiviral.2019.104661
- Primary Citation Related Structures:
6BI8 - PubMed Abstract:
Coronavirus papain-like proteases (PLPs or PLpro), such as the one encoded in the genome of the infectious Middle East Respiratory Syndrome (MERS) virus, have multiple enzymatic activities that promote viral infection. PLpro acts as a protease and processes the large coronavirus polyprotein for virus replication. PLpro also functions as both a deubiquitinating (DUB) and deISGylating (deISG) enzyme and removes ubiquitin (Ub) and interferon-stimulated gene 15 (ISG15) from cellular proteins. Both DUB and deISG activities are implicated in suppressing innate immune responses; however, the precise role of each activity in this process is still unclear due in part to the difficulties in separating each activity. In this study, we determine the first structure of MERS PLpro in complex with the full-length human ISG15 to a resolution of 2.3 Å. This structure and available structures of MERS PLpro-Ub complexes were used as molecular guides to design PLpro mutants that lack either or both DUB/deISG activities. We tested 13 different PLpro mutants for protease, DUB, and deISG activitites using fluorescence-based assays. Results show that we can selectively modulate DUB activity at amino acid positions 1649 and 1653 while mutation of Val1691 or His1652 of PLpro to a positive charged residue completely impairs both DUB/deISG activities. These mutant enzymes will provide new functional tools for delineating the importance of DUB versus deISG activity in virus-infected cells and may serve as potential candidates for attenuating the MERS virus in vivo for modified vaccine design efforts.
- Department of Biological Sciences, Purdue University, West Lafayette, IN, USA.
Organizational Affiliation: