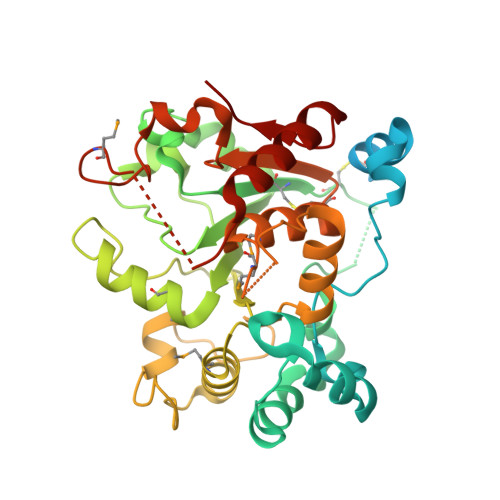
Expression system for structural and functional studies of human glycosylation enzymes.
Moremen, K.W., Ramiah, A., Stuart, M., Steel, J., Meng, L., Forouhar, F., Moniz, H.A., Gahlay, G., Gao, Z., Chapla, D., Wang, S., Yang, J.Y., Prabhakar, P.K., Johnson, R., Rosa, M.D., Geisler, C., Nairn, A.V., Seetharaman, J., Wu, S.C., Tong, L., Gilbert, H.J., LaBaer, J., Jarvis, D.L.(2018) Nat Chem Biol 14: 156-162
- PubMed: 29251719 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.2539
- Primary Citation Related Structures:
6APJ, 6APL - PubMed Abstract:
Vertebrate glycoproteins and glycolipids are synthesized in complex biosynthetic pathways localized predominantly within membrane compartments of the secretory pathway. The enzymes that catalyze these reactions are exquisitely specific, yet few have been extensively characterized because of challenges associated with their recombinant expression as functional products. We used a modular approach to create an expression vector library encoding all known human glycosyltransferases, glycoside hydrolases, and sulfotransferases, as well as other glycan-modifying enzymes. We then expressed the enzymes as secreted catalytic domain fusion proteins in mammalian and insect cell hosts, purified and characterized a subset of the enzymes, and determined the structure of one enzyme, the sialyltransferase ST6GalNAcII. Many enzymes were produced at high yields and at similar levels in both hosts, but individual protein expression levels varied widely. This expression vector library will be a transformative resource for recombinant enzyme production, broadly enabling structure-function studies and expanding applications of these enzymes in glycochemistry and glycobiology.
- Complex Carbohydrate Research Center, University of Georgia, Athens, Georgia, USA.
Organizational Affiliation: