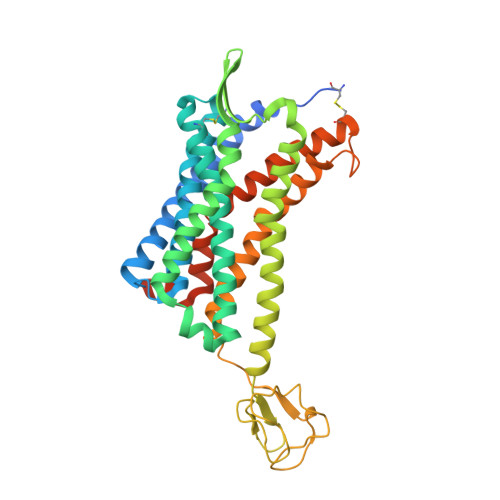
Structure-Based Design of 1-Heteroaryl-1,3-propanediamine Derivatives as a Novel Series of CC-Chemokine Receptor 5 Antagonists.
Peng, P., Chen, H., Zhu, Y., Wang, Z., Li, J., Luo, R.H., Wang, J., Chen, L., Yang, L.M., Jiang, H., Xie, X., Wu, B., Zheng, Y.T., Liu, H.(2018) J Med Chem 61: 9621-9636
- PubMed: 30234300 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01077
- Primary Citation Related Structures:
6AKX, 6AKY - PubMed Abstract:
CC-chemokine receptor 5 (CCR5) is an attractive target for preventing the entry of human immunodeficiency virus 1 (HIV-1) into human host cells. Maraviroc is the only CCR5 antagonist, and it was marketed in 2007. To overcome the shortcomings of maraviroc, structure-based drug design was performed to minimize CYP450 inhibition and to enhance anti-HIV potency and bioavailability. Thirty-four novel 1-heteroaryl-1,3-propanediamine derivatives (1-34) were synthesized, displaying CCR5-antagonist activities in the 2.3-296.4 nM range. Among these, compounds 21 and 34 were the most potent CCR5 antagonists, with excellent in vitro anti-HIV-1 activity, low cytotoxicity, and an acceptable pharmacokinetic profile. Furthermore, the X-ray crystal structures of compounds 21 and 34 bound to CCR5 were determined at 2.8 Å resolution. Compound 34 exhibited no CYP450-inhibition activity at 25 μM, which overcomes the potential drug-drug interaction of maraviroc. Compound 34 represents a promising drug candidate for HIV-infection treatment.
- University of Chinese Academy of Sciences , Number 19A Yuquan Road , Beijing 100049 , China.
Organizational Affiliation: