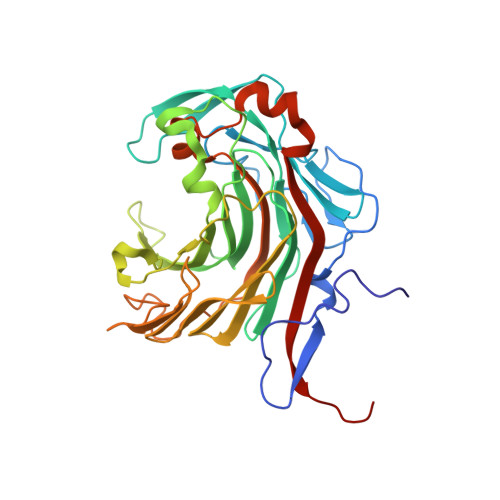
Crystal structure of a neoagarobiose-producing GH16 family beta-agarase from Persicobacter sp. CCB-QB2.
Teh, A.H., Fazli, N.H., Furusawa, G.(2020) Appl Microbiol Biotechnol 104: 633-641
- PubMed: 31784792 Search on PubMed
- DOI: https://doi.org/10.1007/s00253-019-10237-y
- Primary Citation Related Structures:
6AII - PubMed Abstract:
PdAgaC from the marine bacterium Persicobacter sp. CCB-QB2 is a β-agarase belonging to the glycoside hydrolase family 16 (GH16). It is one of only a handful of endo-acting GH16 β-agarases able to degrade agar completely to produce neoagarobiose (NA2). The crystal structure of PdAgaC's catalytic domain, which has one of the highest V max value at 2.9 × 10 3 U/mg, was determined in order to understand its unique mechanism. The catalytic domain is made up of a typical β-jelly roll fold with two additional insertions, and a well-conserved but wider substrate-binding cleft with some minor changes. Among the unique differences, two unconserved residues, Asn226 and Arg286, may potentially contribute additional hydrogen bonds to subsites -1 and +2, respectively, while a third, His185 from one of the additional insertions, may further contribute another bond to subsite +2. These additional hydrogen bonds may probably have enhanced PdAgaC's affinity for short agaro-oligosaccharides such as neoagarotetraose (NA4), rendering it capable of binding NA4 strongly enough for rapid degradation into NA2.
- Centre for Chemical Biology, Universiti Sains Malaysia, 10 Persiaran Bukit Jambul, 11900 Bayan Lepas, Penang, Malaysia. aikhong@usm.my.
Organizational Affiliation: