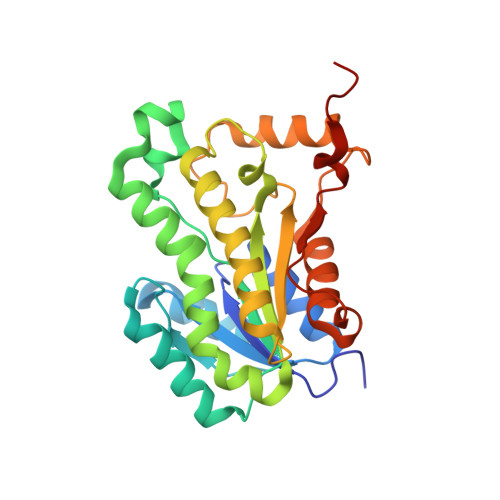
Ternary complex formation of AFN-1252 with Acinetobacter baumannii FabI and NADH: Crystallographic and biochemical studies.
Rao, N.K., Nataraj, V., Ravi, M., Panchariya, L., Palai, K., Talapati, S.R., Lakshminarasimhan, A., Ramachandra, M., Antony, T.(2020) Chem Biol Drug Des 96: 704-713
- PubMed: 32227402 Search on PubMed
- DOI: https://doi.org/10.1111/cbdd.13686
- Primary Citation Related Structures:
6AHE - PubMed Abstract:
Acinetobacter baumannii is an opportunistic Gram-negative bacterial pathogen, associated mostly with hospital-acquired infections. The emergence of drug resistance strains made it necessary to explore new pathways for the development of more effective antibiotics. Enoyl CoA reductase (FabI), a key enzyme in the fatty acid biosynthesis (FAS) pathway, has emerged as a potential target for antibacterial drug development. Earlier reports show that the lead SaFabI inhibitor AFN-1252 can inhibit FabI from other organisms including Escherichia coli and Burkholderia pseudomallei, but with differential potency. In the present work, we show that AFN-1252 is a moderate inhibitor of AbFabI with an IC 50 of 216 nM. AFN-1252 stabilized AbFabI with a 4.2°C increase in the melting temperature (T m ) and, interestingly, the stabilization effect was significantly increased in presence of the cofactor NADH (∆T m = 17°C), suggesting the formation of a ternary complex AbFabI: AFN-1252: NADH. X-ray crystallography studies of AbFabI co-crystalized with AFN-1252 and NADH confirmed the ternary complex formation. The critical interactions of AFN-1252 with AbFabI and NADH identified from the co-crystal structure may facilitate the design and development of new drugs against A. baumannii infections by targeting the FAS pathway.
- Aurigene Discovery Technologies Ltd, Bangalore, India.
Organizational Affiliation: