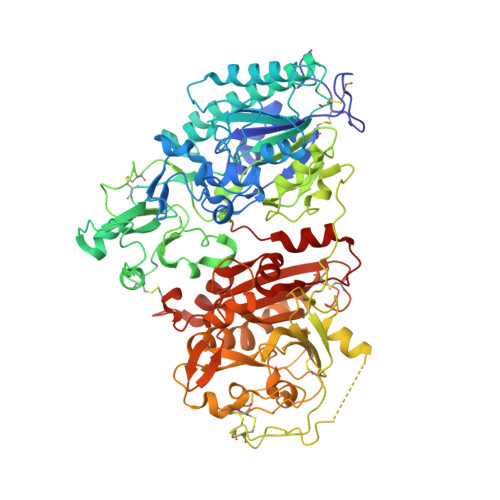
Structural insights into cGAMP degradation by Ecto-nucleotide pyrophosphatase phosphodiesterase 1.
Kato, K., Nishimasu, H., Oikawa, D., Hirano, S., Hirano, H., Kasuya, G., Ishitani, R., Tokunaga, F., Nureki, O.(2018) Nat Commun 9: 4424-4424
- PubMed: 30356045 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-06922-7
- Primary Citation Related Structures:
6AEK, 6AEL - PubMed Abstract:
ENPP1 (Ecto-nucleotide pyrophosphatase phosphodiesterase 1), a type II transmembrane glycoprotein, hydrolyzes ATP to produce AMP and diphosphate, thereby inhibiting bone mineralization. A recent study showed that ENPP1 also preferentially hydrolyzes 2'3'-cGAMP (cyclic GMP-AMP) but not its linkage isomer 3'3'-cGAMP, and negatively regulates the cGAS-STING pathway in the innate immune system. Here, we present the high-resolution crystal structures of ENPP1 in complex with 3'3'-cGAMP and the reaction intermediate pA(3',5')pG. The structures revealed that the adenine and guanine bases of the dinucleotides are recognized by nucleotide- and guanine-pockets, respectively. Furthermore, the structures indicate that 2'3'-cGAMP, but not 3'3'-cGAMP, binds to the active site in a conformation suitable for catalysis, thereby explaining the specific degradation of 2'3'-cGAMP by ENPP1. Our findings provide insights into how ENPP1 hydrolyzes both ATP and cGAMP to participate in the two distinct biological processes.
- Department of Biological Science, Graduate School of Science, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo, 113-0033, Japan.
Organizational Affiliation: