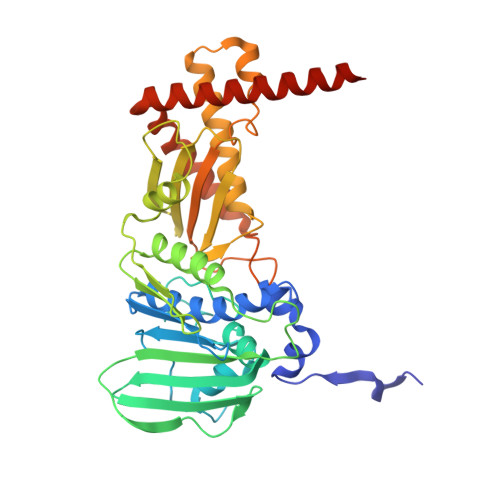
Structural insights into the transient closed conformation and pH dependent ATPase activity of S.Typhi GyraseB N- terminal domain.
Gupta, D., Tiwari, P., Haque, M.A., Sachdeva, E., Hassan, M.I., Ethayathulla, A.S., Kaur, P.(2021) Arch Biochem Biophys 701: 108786-108786
- PubMed: 33548211 Search on PubMed
- DOI: https://doi.org/10.1016/j.abb.2021.108786
- Primary Citation Related Structures:
5ZXM - PubMed Abstract:
DNA Gyrase is a type II topoisomerase that utilizes the energy of ATP hydrolysis for introducing negative supercoils in DNA. The protein comprises two subunits GyrA and GyrB that form a GyrA 2 GyrB 2 heterotetramer. GyrB subunit contains the N-terminal domain (GBNTD) for ATPase activity and the C-terminal domain (GBCTD) for interaction with GyrA and DNA. Earlier structural studies have revealed three different conformational states for GBNTD during ATP hydrolysis defined as open, semi-open, and closed. Here we report, the three-dimensional structure of a new transient closed conformation of GBNTD from Salmonella Typhi (StGBNTD) at 1.94 Å resolution. Based on the structural analysis of this transient closed conformation, we propose the role of protein in the mechanism of ATP hydrolysis. We further explored the effect of pH on ATPase activity and structural stability of the GBNTD using CD and fluorescence spectroscopy at varying pH environment. Kinetic parameters obtained from the ATPase assay were correlated with its secondary and tertiary structure at their respective pH environment. The protein possessed maximum ATPase activity and structural stability at optimum pH 8. At acidic pH, a remarkable decrease in both enzymatic activity and structural stability was observed whereas at alkaline pH there was no significant change. The structural analysis of StGBNTD reveals the role of polar interactions in stabilizing the overall dimeric conformation of the protein.
- Department of Biophysics, All India Institute of Medical Sciences, Ansari Nagar, New Delhi, 110029, India.
Organizational Affiliation: