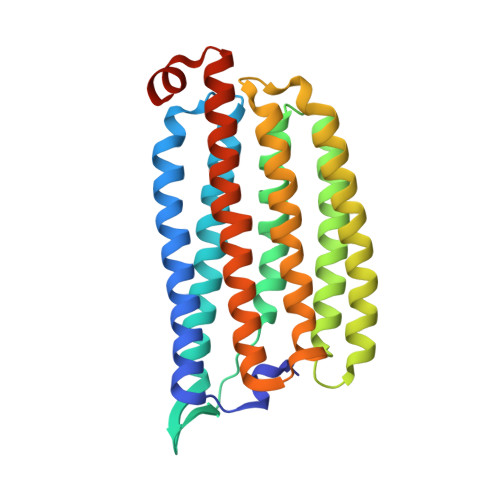
Non-cryogenic structure of a chloride pump provides crucial clues to temperature-dependent channel transport efficiency
Yun, J.H., Li, X., Park, J.H., Wang, Y., Ohki, M., Jin, Z., Lee, W., Park, S.Y., Hu, H., Li, C., Zatsepin, N., Hunter, M.S., Sierra, R.G., Koralek, J., Yoon, C.H., Cho, H.S., Weierstall, U., Tang, L., Liu, H., Lee, W.(2019) J Biological Chem 294: 794-804
- PubMed: 30455349 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA118.004038
- Primary Citation Related Structures:
5ZTK, 5ZTL - PubMed Abstract:
Non-cryogenic protein structures determined at ambient temperature may disclose significant information about protein activity. Chloride-pumping rhodopsin (ClR) exhibits a trend to hyperactivity induced by a change in the photoreaction rate because of a gradual decrease in temperature. Here, to track the structural changes that explain the differences in CIR activity resulting from these temperature changes, we used serial femtosecond crystallography (SFX) with an X-ray free electron laser (XFEL) to determine the non-cryogenic structure of ClR at a resolution of 1.85 Å, and compared this structure with a cryogenic ClR structure obtained with synchrotron X-ray crystallography. The XFEL-derived ClR structure revealed that the all- trans retinal (ATR) region and positions of two coordinated chloride ions slightly differed from those of the synchrotron-derived structure. Moreover, the XFEL structure enabled identification of one additional water molecule forming a hydrogen bond network with a chloride ion. Analysis of the channel cavity and a difference distance matrix plot (DDMP) clearly revealed additional structural differences. B-factor information obtained from the non-cryogenic structure supported a motility change on the residual main and side chains as well as of chloride and water molecules because of temperature effects. Our results indicate that non-cryogenic structures and time-resolved XFEL experiments could contribute to a better understanding of the chloride-pumping mechanism of ClR and other ion pumps.
- From the Department of Biochemistry, College of Life Science & Biotechnology, Yonsei University, Seoul 03722, South Korea.
Organizational Affiliation: