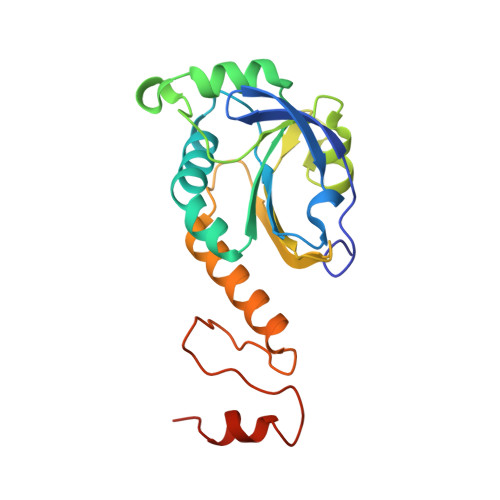
Crystal structure of Arabidopsis thaliana peroxiredoxin A C119S mutant.
Yang, Y., Cai, W., Wang, J., Pan, W., Liu, L., Wang, M., Zhang, M.(2018) Acta Crystallogr F Struct Biol Commun 74: 625-631
- PubMed: 30279313 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X18010920
- Primary Citation Related Structures:
5ZTE - PubMed Abstract:
Peroxiredoxins (Prxs), a large family of antioxidant enzymes, are abundant in all living organisms. Peroxiredoxin A (PrxA) from Arabidopsis thaliana belongs to the typical 2-Cys Prx family and is localized in the chloroplast. This article reports the crystal structure of a PrxA C119S mutant refined to 2.6 Å resolution. The protein exists as a decamer both in the crystal structure and in solution. The structure is in the reduced state suitable for the approach of peroxide, though conformational changes are needed for the resolving process.
- School of Life Sciences, Anhui University, 111 Jiulong Road, Hefei, Anhui 230601, People's Republic of China.
Organizational Affiliation: