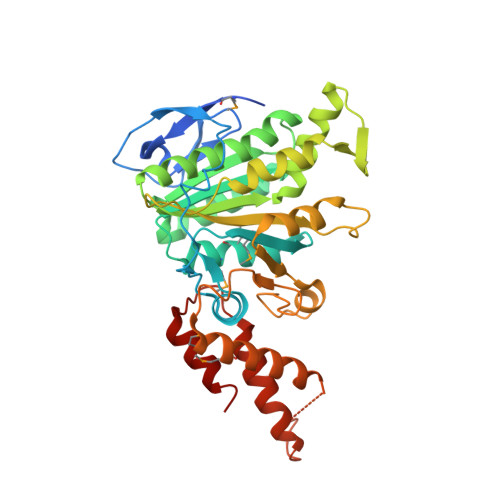
Structural Insight Into Conformational Changes Induced by ATP Binding in a Type III Secretion-Associated ATPase FromShigella flexneri
Gao, X., Mu, Z., Yu, X., Qin, B., Wojdyla, J., Wang, M., Cui, S.(2018) Front Microbiol 9: 1468-1468
- PubMed: 30013545 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fmicb.2018.01468
- Primary Citation Related Structures:
5YBH, 5YBI, 5ZT1 - PubMed Abstract:
Gram-negative bacteria utilize the type III secretion system (T3SS) to inject effector proteins into the host cell cytoplasm, where they subvert cellular functions and assist pathogen invasion. The conserved type III-associated ATPase is critical for the separation of chaperones from effector proteins, the unfolding of effector proteins and translocating them through the narrow channel of the secretion apparatus. However, how ATP hydrolysis is coupled to the mechanical work of the enzyme remains elusive. Herein, we present a complete description of nucleoside triphosphate binding by surface presentation antigens 47 (Spa47) from Shigella flexneri , based on crystal structures containing ATPγS, a catalytic magnesium ion and an ordered water molecule. Combining the crystal structures of Spa47-ATPγS and unliganded Spa47, we propose conformational changes in Spa47 associated with ATP binding, the binding of ATP induces a conformational change of a highly conserved luminal loop, facilitating ATP hydrolysis by the Spa47 ATPase. Additionally, we identified a specific hydrogen bond critical for ATP recognition and demonstrated that, while ATPγS is an ideal analog for probing ATP binding, AMPPNP is a poor ATP mimic. Our findings provide structural insight pertinent for inhibitor design.
- MOH Key Laboratory of Systems Biology of Pathogens, Institute of Pathogen Biology, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing, China.
Organizational Affiliation: