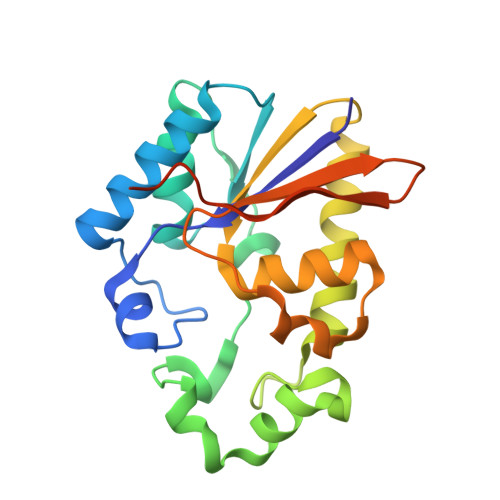
Structural and functional characterisation of phosphoserine phosphatase, that plays critical role in the oxidative stress response in the parasite Entamoeba histolytica.
Kumari, P., Babuta, M., Bhattacharya, A., Gourinath, S.(2019) J Struct Biol 206: 254-266
- PubMed: 30935984 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2019.03.012
- Primary Citation Related Structures:
5ZKK, 5ZR2 - PubMed Abstract:
Amoebiasis is a common parasitic infection in the developing world and is caused by the protist Entameoba histolytica. The proliferation of E. histolytica and its ability to invade epithelial tissues have been shown in several studies to be greatly decreased during oxidative stress. It is therefore not surprising that this amoeba has evolved several mechanisms to evade oxidative stress. Cysteine is thought to be one of the crucial molecules that help in redox defence, and a de novo cysteine biosynthetic pathway involving serine as one of the substrates has been partially elucidated in E. histolytica. Though most of the enzymes of this pathway in E. histolytica have been characterized, phosphoserine phosphatase (EhPSP), a key regulatory enzyme of the serine biosynthetic pathway, has not yet even been identified. In the current work, we identified and characterized EhPSP using various molecular, structural and functional approaches. The crystal structures of native and substrate-bound EhPSP were determined and showed the residues that play a crucial role in its phosphatase activity and substrate binding. Structural and biochemical studies indicated that EhPSP belongs to the histidine phosphatase superfamily. EhPSP-overexpressing amoebic cells were found to be more tolerant to oxidative stress. However, protection during oxidative stress was not seen when a functionally defective mutant was overexpressed. Our results clearly showed that E. histolytica has a functional PSP and that this protein participates in protecting the organism against oxidative stress.
- School of Life Sciences, Jawaharlal Nehru University, New Delhi, India.
Organizational Affiliation: