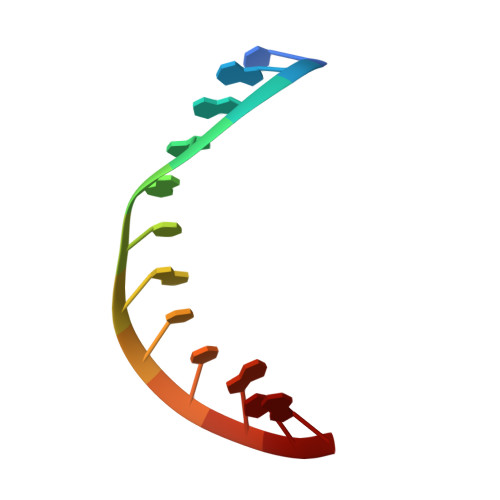
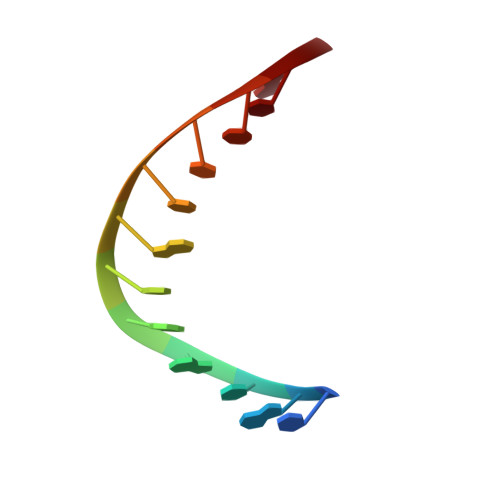
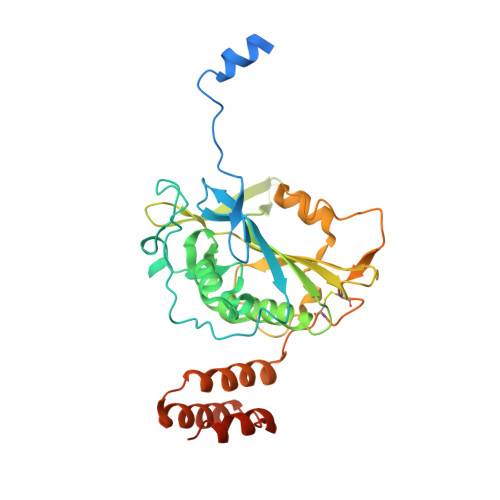
A unique exonuclease ExoG cleaves between RNA and DNA in mitochondrial DNA replication.
Wu, C.C., Lin, J.L.J., Yang-Yen, H.F., Yuan, H.S.(2019) Nucleic Acids Res 47: 5405-5419
- PubMed: 30949702 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz241
- Primary Citation Related Structures:
5ZKI, 5ZKJ, 6IID - PubMed Abstract:
Replication of sufficient mitochondrial DNA (mtDNA) is essential for maintaining mitochondrial functions in mammalian cells. During mtDNA replication, RNA primers must be removed before the nascent circular DNA strands rejoin. This process involves mitochondrial RNase H1, which removes most of the RNA primers but leaves two ribonucleotides attached to the 5' end of nascent DNA. A subsequent 5'-exonuclease is required to remove the residual ribonucleotides, however, it remains unknown if any mitochondrial 5'-exonuclease could remove two RNA nucleotides from a hybrid duplex DNA. Here, we report that human mitochondrial Exonuclease G (ExoG) may participate in this particular process by efficiently cleaving at RNA-DNA junctions to remove the 5'-end RNA dinucleotide in an RNA/DNA hybrid duplex. Crystal structures of human ExoG bound respectively with DNA, RNA/DNA hybrid and RNA-DNA chimeric duplexes uncover the underlying structural mechanism of how ExoG specifically recognizes and cleaves at RNA-DNA junctions of a hybrid duplex with an A-form conformation. This study hence establishes the molecular basis of ExoG functioning as a unique 5'-exonuclease to mediate the flap-independent RNA primer removal process during mtDNA replication to maintain mitochondrial genome integrity.
- Institute of Molecular Biology, Academia Sinica, Taipei, Taiwan 11529, ROC.
Organizational Affiliation: