Crystal structures of unlinked full length NS3 from Dengue virus provide insights into dynamics of protease domain
Phoo, W.W., Sahili, A.E., Zhang, Z.Z., Chen, M.W., Vasudevan, S., Lescar, J., Luo, D.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Genome polyprotein | A [auth B] | 618 | dengue virus type 4 | Mutation(s): 1 |  |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | F8TEL4 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Genome polyprotein | B [auth C] | 47 | dengue virus type 4 | Mutation(s): 0 |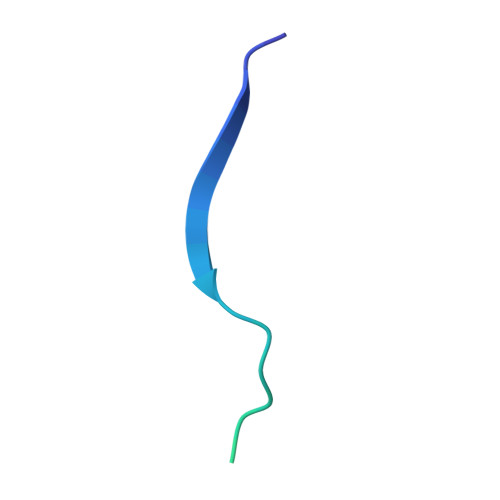  |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q2YHE8 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GOL Download:Ideal Coordinates CCD File | C [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 52.78 | α = 90 |
| b = 86.81 | β = 93.07 |
| c = 75.9 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| iMOSFLM | data reduction |
| SCALA | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| NTU | Singapore | START UP GRANT |
| NMRC | Singapore | CBRG14May05 |