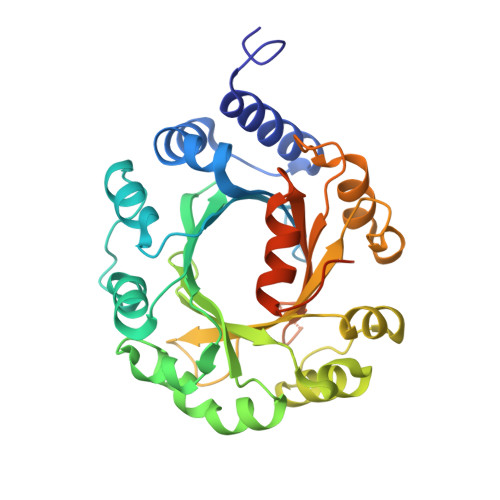
Structural insight into a novel indole prenyltransferase in hapalindole-type alkaloid biosynthesis.
Wang, J., Chen, C.C., Yang, Y., Liu, W., Ko, T.P., Shang, N., Hu, X., Xie, Y., Huang, J.W., Zhang, Y., Guo, R.T.(2018) Biochem Biophys Res Commun 495: 1782-1788
- PubMed: 29229390 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2017.12.039
- Primary Citation Related Structures:
5YNT, 5YNU, 5YNV, 5YNW - PubMed Abstract:
FamD1 is a novel CloQ/NphB-family indole prenyltransferase which involves in hapalindole-type alkaloid biosynthesis. Here the native FamD1 structure and three protein-ligand complexes are analyzed to investigate the molecular basis of substrate binding and catalysis. FamD1 adopts a typical ABBA architecture of aromatic prenyltransferase, in which the substrate-binding chamber is found in the central β-barrel. The indole-containing acceptor substrate is bound adjacent to the prenyl donor. Based on the complex structures, a catalytic mechanism of FamD1 is proposed. Functional implications on the sister enzyme FamD2 are also discussed.
- Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, Tianjin 300308, China.
Organizational Affiliation: