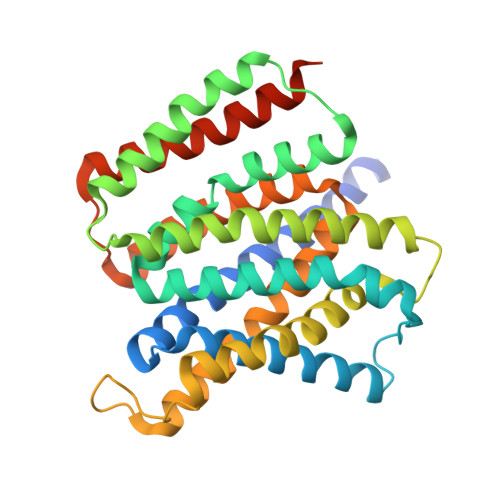
Structure of the triose-phosphate/phosphate translocator reveals the basis of substrate specificity
Lee, Y., Nishizawa, T., Takemoto, M., Kumazaki, K., Yamashita, K., Hirata, K., Minoda, A., Nagatoishi, S., Tsumoto, K., Ishitani, R., Nureki, O.(2017) Nat Plants 3: 825-832
- PubMed: 28970497 Search on PubMed
- DOI: https://doi.org/10.1038/s41477-017-0022-8
- Primary Citation Related Structures:
5Y78, 5Y79 - PubMed Abstract:
The triose-phosphate/phosphate translocator (TPT) catalyses the strict 1:1 exchange of triose-phosphate, 3-phosphoglycerate and inorganic phosphate across the chloroplast envelope, and plays crucial roles in photosynthesis. Despite rigorous study for more than 40 years, the molecular mechanism of TPT is poorly understood because of the lack of structural information. Here we report crystal structures of TPT bound to two different substrates, 3-phosphoglycerate and inorganic phosphate, in occluded conformations. The structures reveal that TPT adopts a 10-transmembrane drug/metabolite transporter fold. Both substrates are bound within the same central pocket, where conserved lysine, arginine and tyrosine residues recognize the shared phosphate group. A structural comparison with the outward-open conformation of the bacterial drug/metabolite transporter suggests a rocker-switch motion of helix bundles, and molecular dynamics simulations support a model in which this rocker-switch motion is tightly coupled to the substrate binding, to ensure strict 1:1 exchange. These results reveal the unique mechanism of sugar phosphate/phosphate exchange by TPT.
- Department of Biological Sciences, Graduate School of Science, University of Tokyo, 2-11-16 Yayoi, Bunkyo-ku, Tokyo, 113-0032, Japan.
Organizational Affiliation: