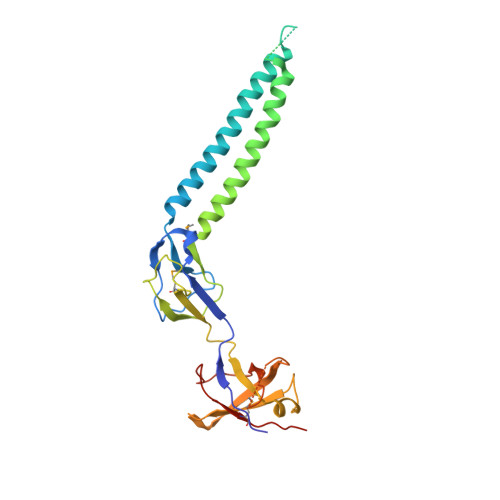
Structure of a MacAB-like efflux pump from Streptococcus pneumoniae.
Yang, H.B., Hou, W.T., Cheng, M.T., Jiang, Y.L., Chen, Y., Zhou, C.Z.(2018) Nat Commun 9: 196-196
- PubMed: 29335499 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02741-4
- Primary Citation Related Structures:
5XU0, 5XU1 - PubMed Abstract:
The spr0693-spr0694-spr0695 operon of Streptococcus pneumoniae encodes a putative ATP-binding cassette (ABC)-type efflux pump involved in the resistance of antibiotics and antimicrobial peptides. Here we report the crystal structures of Spr0694-0695 at 3.3 Å and Spr0693 at 3.0 Å resolution, revealing a MacAB-like efflux pump. The dimeric Spr0694-0695 adopts a non-canonical fold of ABC transporter, the transmembrane domain of which consists of eight tightly packed transmembrane helices with an insertion of extracellular domain between the first and second helices, whereas Spr0693 forms a nanotube channel docked onto the ABC transporter. Structural analyses combined with ATPase activity and antimicrobial susceptibility assays, enable us to propose a putative substrate-entrance tunnel with a lateral access controlled by a guard helix. Altogether, our findings provide structural insights and putative transport mechanism of a MacAB-like efflux pump in Gram-positive bacteria.
- Hefei National Laboratory for Physical Sciences at the Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, 230027, China.
Organizational Affiliation: