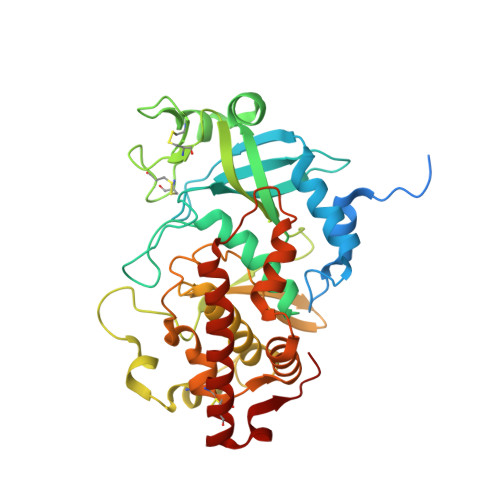
Structure and evolution of the Fam20 kinases
Zhang, H., Zhu, Q., Cui, J., Wang, Y., Chen, M.J., Guo, X., Tagliabracci, V.S., Dixon, J.E., Xiao, J.(2018) Nat Commun 9: 1218-1218
- PubMed: 29572475 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-03615-z
- Primary Citation Related Structures:
5XOM, 5XOO, 5YH0, 5YH2, 5YH3 - PubMed Abstract:
The Fam20 proteins are novel kinases that phosphorylate secreted proteins and proteoglycans. Fam20C phosphorylates hundreds of secreted proteins and is activated by the pseudokinase Fam20A. Fam20B phosphorylates a xylose residue to regulate proteoglycan synthesis. Despite these wide-ranging and important functions, the molecular and structural basis for the regulation and substrate specificity of these kinases are unknown. Here we report molecular characterizations of all three Fam20 kinases, and show that Fam20C is activated by the formation of an evolutionarily conserved homodimer or heterodimer with Fam20A. Fam20B has a unique active site for recognizing Galβ1-4Xylβ1, the initiator disaccharide within the tetrasaccharide linker region of proteoglycans. We further show that in animals the monomeric Fam20B preceded the appearance of the dimeric Fam20C, and the dimerization trait of Fam20C emerged concomitantly with a change in substrate specificity. Our results provide comprehensive structural, biochemical, and evolutionary insights into the function of the Fam20 kinases.
- The State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking-Tsinghua Center for Life Sciences, Peking University, 100871, Beijing, China.
Organizational Affiliation: