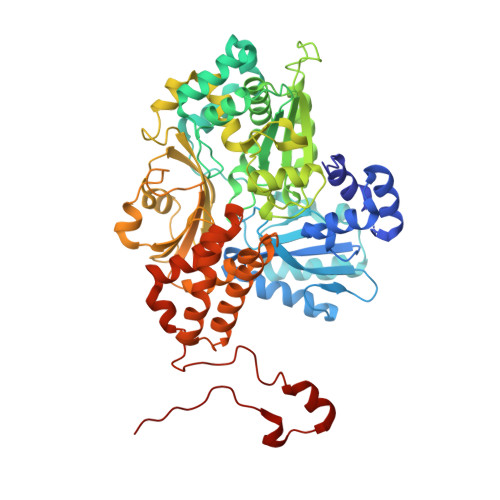
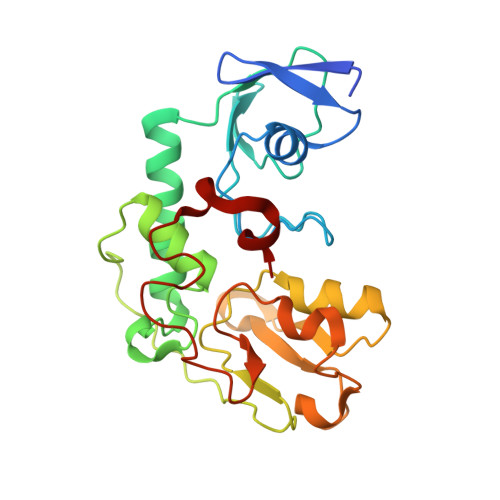
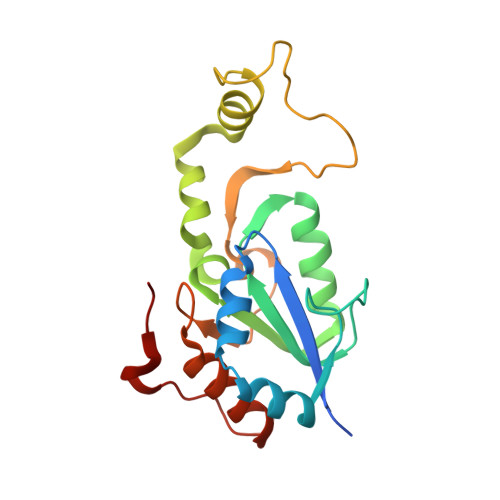
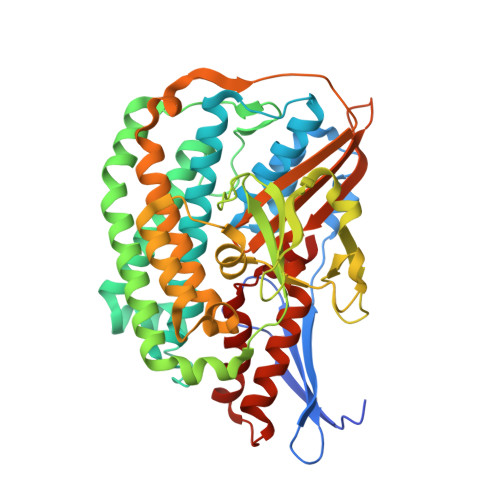
Structural basis of the redox switches in the NAD(+)-reducing soluble [NiFe]-hydrogenase
Shomura, Y., Taketa, M., Nakashima, H., Tai, H., Nakagawa, H., Ikeda, Y., Ishii, M., Igarashi, Y., Nishihara, H., Yoon, K.S., Ogo, S., Hirota, S., Higuchi, Y.(2017) Science 357: 928-932
- PubMed: 28860386 Search on PubMed
- DOI: https://doi.org/10.1126/science.aan4497
- Primary Citation Related Structures:
5XF9, 5XFA - PubMed Abstract:
NAD + (oxidized form of NAD:nicotinamide adenine dinucleotide)-reducing soluble [NiFe]-hydrogenase (SH) is phylogenetically related to NADH (reduced form of NAD + ):quinone oxidoreductase (complex I), but the geometrical arrangements of the subunits and Fe-S clusters are unclear. Here, we describe the crystal structures of SH in the oxidized and reduced states. The cluster arrangement is similar to that of complex I, but the subunits orientation is not, which supports the hypothesis that subunits evolved as prebuilt modules. The oxidized active site includes a six-coordinate Ni, which is unprecedented for hydrogenases, whose coordination geometry would prevent O 2 from approaching. In the reduced state showing the normal active site structure without a physiological electron acceptor, the flavin mononucleotide cofactor is dissociated, which may be caused by the oxidation state change of nearby Fe-S clusters and may suppress production of reactive oxygen species.
- Institute of Quantum Beam Science, Graduate School of Science and Engineering, Ibaraki University, 4-12-1 Nakanarusawa, Hitachi, Ibaraki 316-8511, Japan. hig@sci.u-hyogo.ac.jp yasuhito.shomura.s@vc.ibaraki.ac.jp.
Organizational Affiliation: