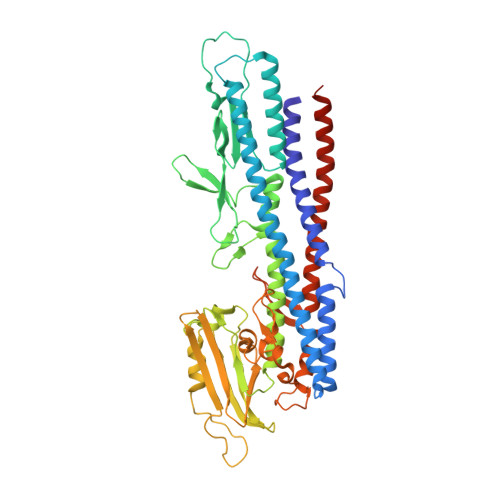
Structure of FlgK reveals the divergence of the bacterial Hook-Filament Junction of Campylobacter
Bulieris, P.V., Shaikh, N.H., Freddolino, P.L., Samatey, F.A.(2017) Sci Rep 7: 15743-15743
- PubMed: 29147015 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-15837-0
- Primary Citation Related Structures:
5XBJ - PubMed Abstract:
Evolution of a nano-machine consisting of multiple parts, each with a specific function, is a complex process. A change in one part should eventually result in changes in other parts, if the overall function is to be conserved. In bacterial flagella, the filament and the hook have distinct functions and their respective proteins, FliC and FlgE, have different three-dimensional structures. The filament functions as a helical propeller and the hook as a flexible universal joint. Two proteins, FlgK and FlgL, assure a smooth connectivity between the hook and the filament. Here we show that, in Campylobacter, the 3D structure of FlgK differs from that of its orthologs in Salmonella and Burkholderia, whose structures have previously been solved. Docking the model of the FlgK junction onto the structure of the Campylobacter hook provides some clues about its divergence. These data suggest how evolutionary pressure to adapt to structural constraints, due to the structure of Campylobacter hook, causes divergence of one element of a supra-molecular complex in order to maintain the function of the entire flagellar assembly.
- Trans-membrane Trafficking Unit, Okinawa Institute of Science and Technology Graduate University, 1919-1 Tancha, Onna, Kunigami, Okinawa, 904-0495, Japan.
Organizational Affiliation: