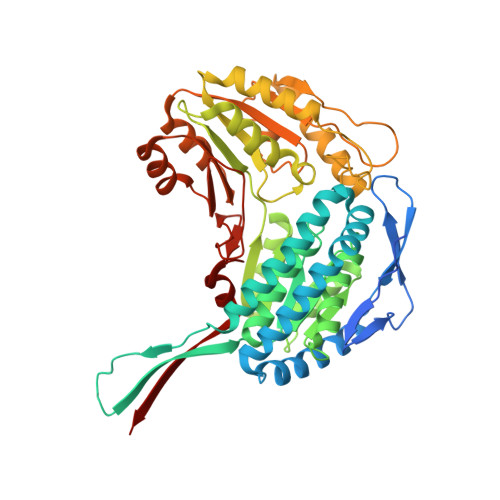
Structural insights into the production of 3-hydroxypropionic acid by aldehyde dehydrogenase from Azospirillum brasilense.
Son, H.F., Park, S., Yoo, T.H., Jung, G.Y., Kim, K.J.(2017) Sci Rep 7: 46005-46005
- PubMed: 28393833 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep46005
- Primary Citation Related Structures:
5X5T, 5X5U - PubMed Abstract:
3-Hydroxypropionic acid (3-HP) is an important platform chemical to be converted to acrylic acid and acrylamide. Aldehyde dehydrogenase (ALDH), an enzyme that catalyzes the reaction of 3-hydroxypropionaldehyde (3-HPA) to 3-HP, determines 3-HP production rate during the conversion of glycerol to 3-HP. To elucidate molecular mechanism of 3-HP production, we determined the first crystal structure of a 3-HP producing ALDH, α-ketoglutarate-semialdehyde dehydrogenase from Azospirillum basilensis (AbKGSADH), in its apo-form and in complex with NAD + . Although showing an overall structure similar to other ALDHs, the AbKGSADH enzyme had an optimal substrate binding site for accepting 3-HPA as a substrate. Molecular docking simulation of 3-HPA into the AbKGSADH structure revealed that the residues Asn159, Gln160 and Arg163 stabilize the aldehyde- and the hydroxyl-groups of 3-HPA through hydrogen bonds, and several hydrophobic residues, such as Phe156, Val286, Ile288, and Phe450, provide the optimal size and shape for 3-HPA binding. We also compared AbKGSADH with other reported 3-HP producing ALDHs for the crucial amino acid residues for enzyme catalysis and substrate binding, which provides structural implications on how these enzymes utilize 3-HPA as a substrate.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, Daehak-ro 80, Buk-ku, Daegu 702-701, Korea.
Organizational Affiliation: