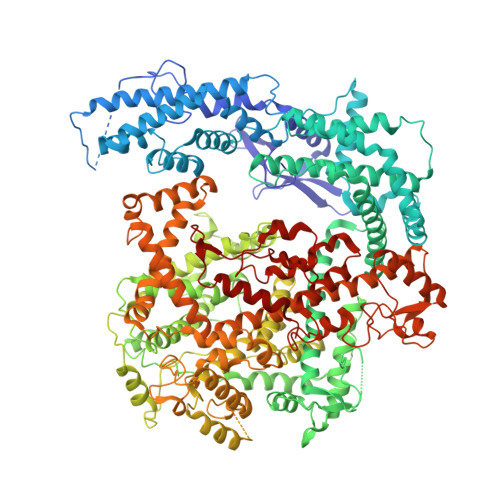
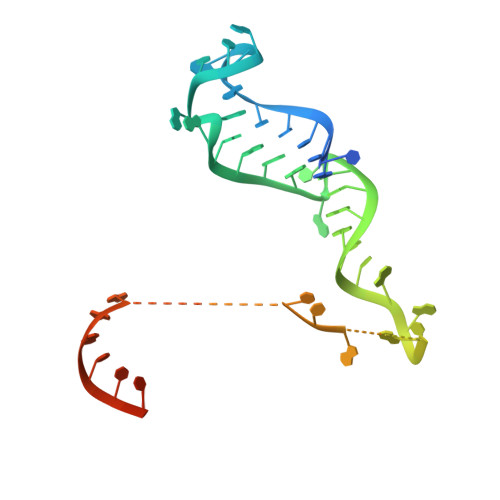
Two Distant Catalytic Sites Are Responsible for C2c2 RNase Activities
Liu, L., Li, X., Wang, J., Wang, M., Chen, P., Yin, M., Li, J., Sheng, G., Wang, Y.(2017) Cell 168: 121-134.e12
- PubMed: 28086085 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2016.12.031
- Primary Citation Related Structures:
5WTJ, 5WTK - PubMed Abstract:
C2c2, the effector of type VI CRISPR-Cas systems, has two RNase activities-one for cutting its RNA target and the other for processing the CRISPR RNA (crRNA). Here, we report the structures of Leptotrichia shahii C2c2 in its crRNA-free and crRNA-bound states. While C2c2 has a bilobed structure reminiscent of all other Class 2 effectors, it also exhibits different structural characteristics. It contains the REC lobe with a Helical-1 domain and the NUC lobe with two HEPN domains. The two RNase catalytic pockets responsible for cleaving pre-crRNA and target RNA are independently located on Helical-1 and HEPN domains, respectively. crRNA binding induces significant conformational changes that are likely to stabilize crRNA binding and facilitate target RNA recognition. These structures provide important insights into the molecular mechanism of dual RNase activities of C2c2 and establish a framework for its future engineering as a RNA editing tool.
- Key Laboratory of RNA Biology, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China.
Organizational Affiliation: