Identification of Near-Pan-neutralizing Antibodies against HIV-1 by Deconvolution of Plasma Humoral Responses.
Sajadi, M.M., Dashti, A., Rikhtegaran Tehrani, Z., Tolbert, W.D., Seaman, M.S., Ouyang, X., Gohain, N., Pazgier, M., Kim, D., Cavet, G., Yared, J., Redfield, R.R., Lewis, G.K., DeVico, A.L.(2018) Cell 173: 1783-1795.e14
- PubMed: 29731169 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2018.03.061
- Primary Citation Related Structures:
5WB9, 6BCK - PubMed Abstract:
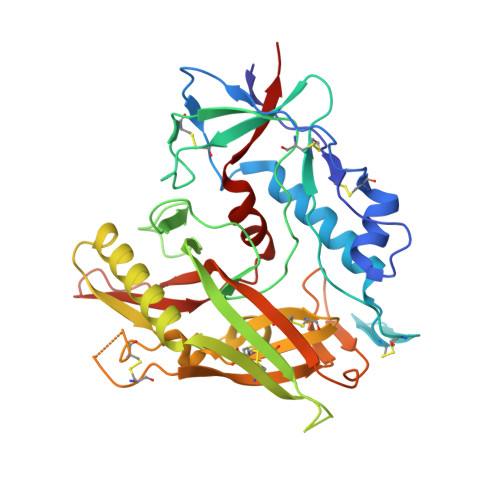
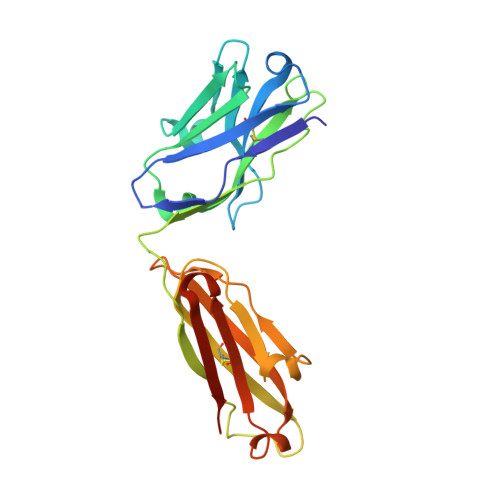
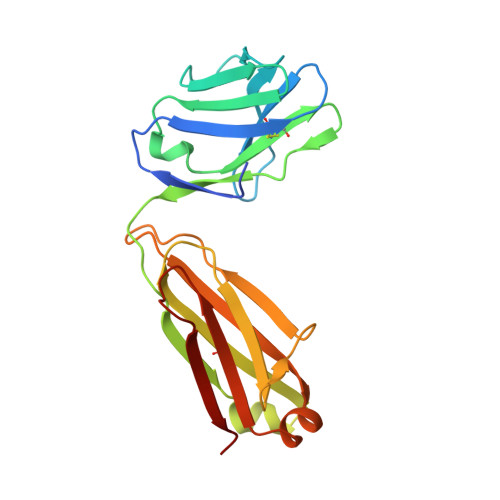
Anti-HIV-1 envelope broadly neutralizing monoclonal antibodies (bNAbs) isolated from memory B cells may not fully represent HIV-1-neutralizing profiles measured in plasma. Accordingly, we characterized near-pan-neutralizing antibodies extracted directly from the plasma of two "elite neutralizers." Circulating anti-gp120 polyclonal antibodies were deconvoluted using proteomics to guide lineage analysis of bone marrow plasma cells. In both subjects, a single lineage of anti-CD4-binding site (CD4bs) antibodies explained the plasma-neutralizing activity. Importantly, members of these lineages potently neutralized 89%-100% of a multi-tier 117 pseudovirus panel, closely matching the specificity and breadth of the circulating antibodies. X-ray crystallographic analysis of one monoclonal, N49P7, suggested a unique ability to bypass the CD4bs Phe43 cavity, while reaching deep into highly conserved residues of Layer 3 of the gp120 inner domain, likely explaining its extreme potency and breadth. Further direct analyses of plasma anti-HIV-1 bNAbs should provide new insights for developing antibody-based antiviral agents and vaccines.
- Divisions of Vaccine Research and Clinical Care and Research, Institute of Human Virology, University of Maryland School of Medicine, Baltimore, MD 21201, USA; Department of Medicine, Baltimore VA Medical Center, Baltimore, MD 21201, USA; Department of Medicine, University of Maryland School of Medicine, Baltimore, MD 21201, USA. Electronic address: msajadi@ihv.umaryland.edu.
Organizational Affiliation: