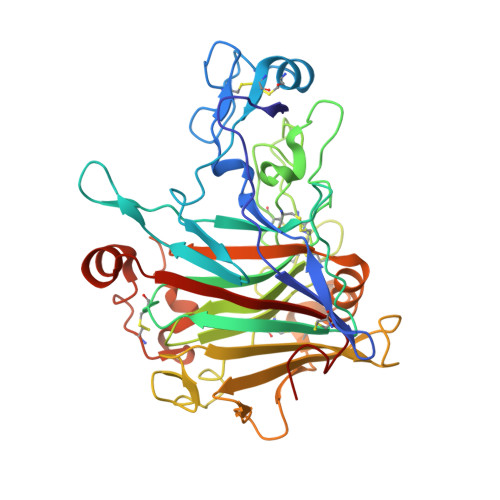
Structure and dynamics of Trichoderma harzianum Cel7B suggest molecular architecture adaptations required for a wide spectrum of activities on plant cell wall polysaccharides.
Sonoda, M.T., Godoy, A.S., Pellegrini, V.O.A., Kadowaki, M.A.S., Nascimento, A.S., Polikarpov, I.(2019) Biochim Biophys Acta Gen Subj 1863: 1015-1026
- PubMed: 30898558 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbagen.2019.03.013
- Primary Citation Related Structures:
5W0A - PubMed Abstract:
Cellulases from glycoside hydrolase family 7 (GH7) play crucial roles in plant lignocellulose deconstruction by fungi, but structural information available for GH7 fungal endoglucanases is limited when compared to the number of known sequences in the family. Here, we report the X-ray structure of the glycosylated catalytic domain (CD) of Trichoderma harzianum endoglucanase, ThCel7B, solved and refined at 2.9 Å resolution. Additionally, our extensive molecular dynamics simulations of this enzyme in complex with a variety of oligosaccharides provide a better understanding of its promiscuous hydrolytic activities on plant cell wall polysaccharides. The simulations demonstrate the importance of the hydrogen bond between substrate O2 hydroxyl in the subsite -1 and a side chain of catalytic Glu196 which renders ThCel7B capable to catalytically cleave cello and xylooligosaccharides, but not mannooligosaccharides. Moreover, detailed structural analyses and MD simulations revealed an additional binding pocket, suitable for accommodation of oligosaccharide decorations and/or substrates with mixed glycoside bonds that abuts onto the binding cleft close to subsite +2.
- Departamento de Física, Instituto de Ciências Exatas, Naturais e Educação, Universidade Federal do Triângulo Mineiro, Av. Frei Paulino, 30 - Bairro Abadia Uberaba, MG 38025-180, Brazil.
Organizational Affiliation: