Tuning the Cavity Size and Chirality of Self-Assembling 3D DNA Crystals.
Simmons, C.R., Zhang, F., MacCulloch, T., Fahmi, N., Stephanopoulos, N., Liu, Y., Seeman, N.C., Yan, H.(2017) J Am Chem Soc 139: 11254-11260
- PubMed: 28731332 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.7b06485
- Primary Citation Related Structures:
5VY6, 5VY7 - PubMed Abstract:
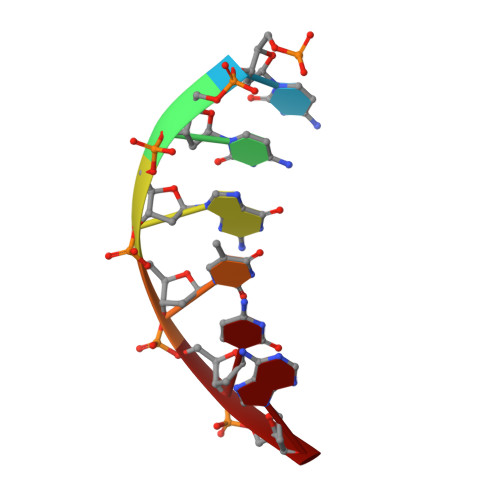
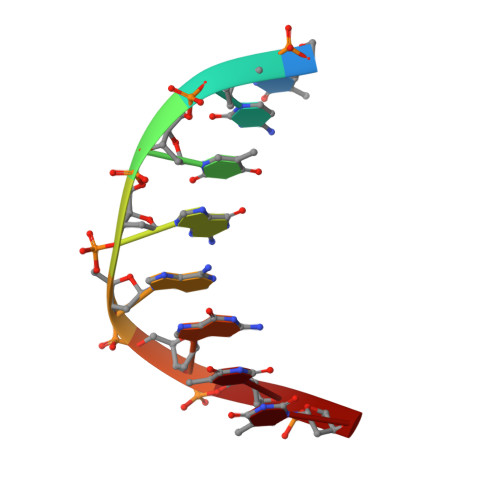
The foundational goal of structural DNA nanotechnology-the field that uses oligonucleotides as a molecular building block for the programmable self-assembly of nanostructured systems-was to use DNA to construct three-dimensional (3D) lattices for solving macromolecular structures. The programmable nature of DNA makes it an ideal system for rationally constructing self-assembled crystals and immobilizing guest molecules in a repeating 3D array through their specific stereospatial interactions with the scaffold. In this work, we have extended a previously described motif (4 × 5) by expanding the structure to a system that links four double-helical layers; we use a central weaving oligonucleotide containing a sequence of four six-base repeats (4 × 6), forming a matrix of layers that are organized and dictated by a series of Holliday junctions. In addition, we have assembled mirror image crystals (l-DNA) with the identical sequence that are completely resistant to nucleases. Bromine and selenium derivatives were obtained for the l- and d-DNA forms, respectively, allowing phase determination for both forms and solution of the resulting structures to 3.0 and 3.05 Å resolution. Both right- and left-handed forms crystallized in the trigonal space groups with mirror image 3-fold helical screw axes P3 2 and P3 1 for each motif, respectively. The structures reveal a highly organized array of discrete and well-defined cavities that are suitable for hosting guest molecules and allow us to dictate a priori the assembly of guest-DNA conjugates with a specified crystalline hand.
- Department of Chemistry, New York University , New York, New York 10003, United States.
Organizational Affiliation: